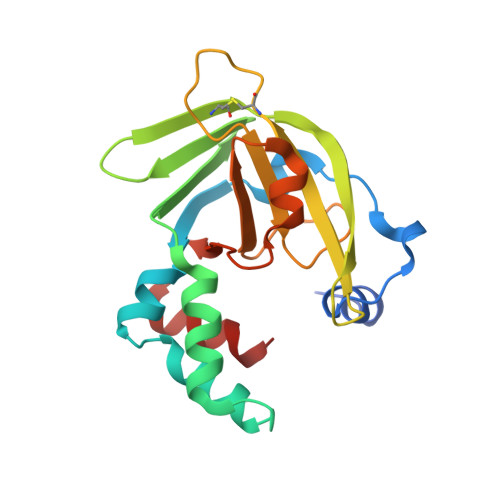
Yoda from Escherichia Coli is a Metal-Binding, Lipocalin-Like Protein
David, G., Blondeau, K., Schiltz, M., Penel, S., Lewit-Bentley, A.(2003) J Biological Chem 278: 43728
- PubMed: 12909634 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M304484200
- Primary Citation Related Structures:
1OEE, 1OEJ, 1OEK - PubMed Abstract:
We have determined the crystal structure of YodA, an Escherichia coli protein of unknown function. YodA had been identified under conditions of cadmium stress, and we confirm that it binds metals such as cadmium and zinc. We have also found nickel bound in one of the crystal forms. YodA is composed of two domains: a main lipocalin/calycin-like domain and a helical domain. The principal metal-binding site lies on one side of the calycin domain, thus making YodA the first metal-binding lipocalin known. Our experiments suggest that YodA expression may be part of a more general stress response. From sequence analogy with the C-terminal domain of a metal-binding receptor of a member of bacterial ATP-binding cassette transporters, we propose a three-dimensional model for this receptor and suggest that YodA may have a receptor-type partner in E. coli.
- Laboratoire pour l'Utilisation du Rayonnement Electromagnétique, CNRS, CEA, MdR, BP. 34, 91898 Orsay Cedex, France.
Organizational Affiliation: