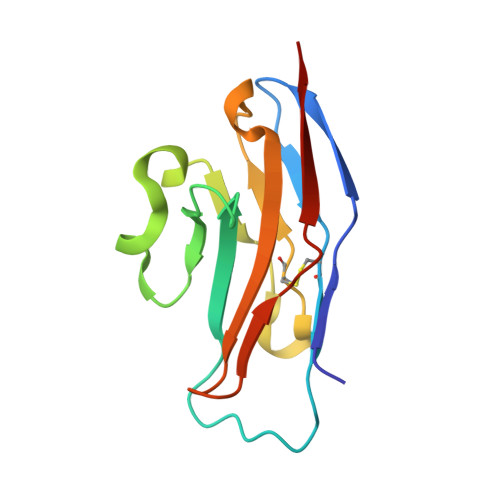
Structure-Guided Design of Sialic Acid-Based Siglec Inhibitors and Crystallographic Analysis in Complex with Sialoadhesin
Zaccai, N.R., Maenaka, K., Maenaka, T., Crocker, P.R., Brossmer, R., Kelm, S., Jones, E.Y.(2003) Structure 11: 557
- PubMed: 12737821 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(03)00073-x
- Primary Citation Related Structures:
1OD7, 1OD9, 1ODA - PubMed Abstract:
The Siglec family of receptors mediates cell surface interactions through recognition of sialylated glycoconjugates. The crystal structure of the N-terminal immunoglobulin-like domain of the Siglec sialoadhesin (SnD1) in complex with 2,3-sialyllactose has informed the design of sialic acid analogs (sialosides) that bind Siglecs with significantly enhanced affinities and specificities. Binding assays against sialoadhesin (Sn; Siglec-1), CD22 (Siglec-2), and MAG (Siglec-4) show a 10- to 300-fold reduction in IC(50) values (relative to methyl-alpha-Neu5Ac) for three sialosides bearing aromatic group modifications of the glycerol side chain: Me-alpha-9-N-benzoyl-amino-9-deoxy-Neu5Ac (BENZ), Me-alpha-9-N-(naphthyl-2-carbonyl)-amino-9-deoxy-Neu5Ac (NAP), and Me-alpha-9-N-(biphenyl-4-carbonyl)-amino-9-deoxy-Neu5Ac (BIP). Crystal structures of these sialosides in complex with SnD1 suggest explanations for the differences in specificity and affinity, providing further ideas for compound design of physiological and potentially therapeutic relevance.
- CR-UK Receptor Structure Research Group, Division of Structural Biology, The Henry Wellcome Building for Genomic Medicine, Roosevelt Drive, Headington, Oxford OX3 7BN, United Kingdom.
Organizational Affiliation: