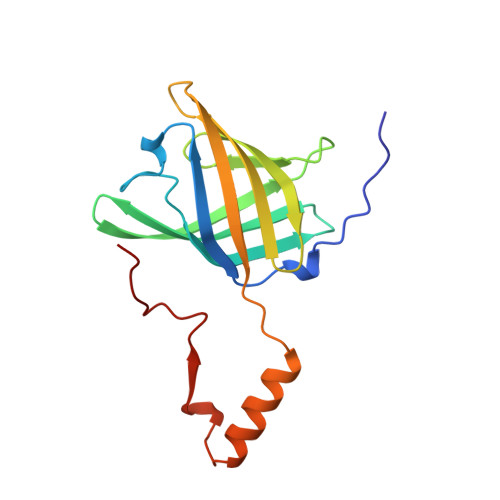
Domain swapping creates a third putative combining site in bovine odorant binding protein dimer
Tegoni, M., Ramoni, R., Bignetti, E., Spinelli, S., Cambillau, C.(1996) Nat Struct Biol 3: 863-867
- PubMed: 8836103 Search on PubMed
- DOI: https://doi.org/10.1038/nsb1096-863
- Primary Citation Related Structures:
1OBP - PubMed Abstract:
In mammals, odorant binding proteins may play an important role in the transport of odors towards specific olfactory receptors on sensory neurones across the aqueous compartment of the nasal mucus. We have solved the X-ray structure of such a transport protein, bovine odorant binding protein (OBP) at 2.0 A resolution. The beta-barrel of OBP is similar to that of lipocalins, but OBP dimer association results from domain swapping, an observation unique among the lipocalins. The alpha-helix of each monomer stacks against the beta-barrel of the other monomer. Contrary to previous reports, each monomer has an internal buried cavity which could accommodate a naturally occurring molecule. Besides this cavity, an open cavity is located at the dimer interface. Data in solution suggest that this central cavity may be a binding site created by domain swapping.
- UPR9039-CNRS, IFR1, Marseille, France.
Organizational Affiliation: