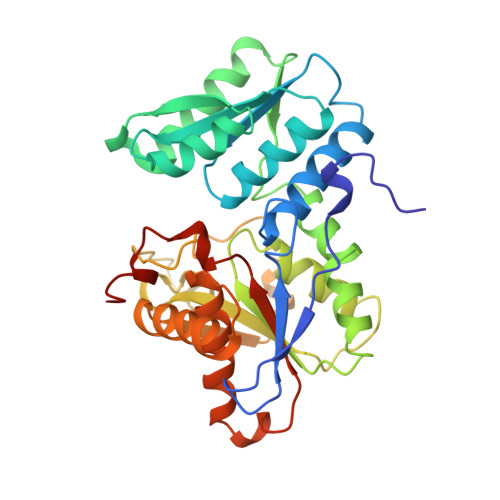
Three-dimensional structure of O-acetylserine sulfhydrylase from Salmonella typhimurium.
Burkhard, P., Rao, G.S., Hohenester, E., Schnackerz, K.D., Cook, P.F., Jansonius, J.N.(1998) J Mol Biology 283: 121-133
- PubMed: 9761678 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1998.2037
- Primary Citation Related Structures:
1OAS - PubMed Abstract:
The last step in cysteine biosynthesis in enteric bacteria is catalyzed by the pyridoxal 5'-phosphate-dependent enzyme O-acetylserine sulfhydrylase. Here we report the crystal structure at 2.2 A resolution of the A-isozyme of O-acetylserine sulfhydrylase isolated from Salmonella typhimurium. O-acetylserine sulfhydrylase shares the same fold with tryptophan synthase-beta from Salmonella typhimurium but the sequence identity level is below 20%. There are some major structural differences: the loops providing the interface to the alpha-subunit in tryptophan synthase-beta and two surface helices of tryptophan synthase-beta are missing in O-acetylserine sulfhydrylase. The hydrophobic channel for indole transport from the alpha to the beta active site of tryptophan synthase-beta is, not unexpectedly, also absent in O-acetylserine sulfhydrylase. The dimer interface, on the other hand, is more or less conserved in the two enzymes. The active site cleft of O-acetylserine sulfhydrylase is wider and therefore more exposed to the solvent. A possible binding site for the substrate O-acetylserine is discussed.
- Department of Structural Biology, Biozentrum, University of Basel, Klingelbergstrasse70, Basel, CH-4056, Switzerland.
Organizational Affiliation: