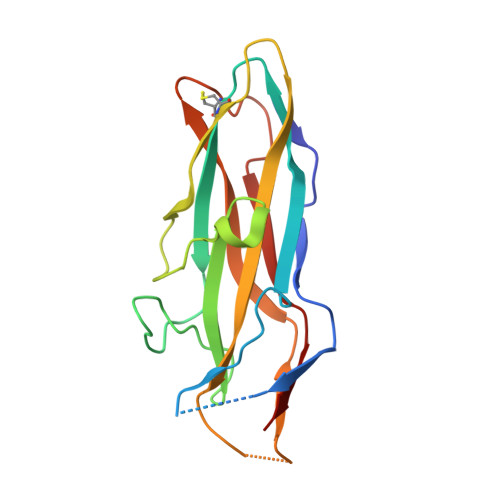
The Fimbrial Adhesin F17-G of Enterotoxigenic Escherichia Coli Has an Immunoglobulin-Like Lectin Domain that Binds N-Acetylglucosamine
Buts, L., Bouckaert, J., De Genst, E., Loris, R., Oscarson, S., Lahmann, M., Messens, J., Brosens, E., Wyns, L., De Greve, H.(2003) Mol Microbiol 49: 705
- PubMed: 12864853 Search on PubMed
- DOI: https://doi.org/10.1046/j.1365-2958.2003.03600.x
- Primary Citation Related Structures:
1O9V, 1O9W, 1O9Z - PubMed Abstract:
The F17-G adhesin at the tip of flexible F17 fimbriae of enterotoxigenic Escherichia coli mediates binding to N-acetyl-beta-D-glucosamine-presenting receptors on the microvilli of the intestinal epithelium of ruminants. We report the 1.7 A resolution crystal structure of the lectin domain of F17-G, both free and in complex with N-acetylglucosamine. The monosaccharide is bound on the side of the ellipsoid-shaped protein in a conserved site around which all natural variations of F17-G are clustered. A model is proposed for the interaction between F17-fimbriated E. coli and microvilli with enhanced affinity compared with the binding constant we determined for F17-G binding to N-acetylglucosamine (0.85 mM-1). Unexpectedly, the F17-G structure reveals that the lectin domains of the F17-G, PapGII and FimH fimbrial adhesins all share the immunoglobulin-like fold of the structural components (pilins) of their fimbriae, despite lack of any sequence identity. Fold comparisons with pilin and chaperone structures of the chaperone/usher pathway highlight the central role of the C-terminal beta-strand G of the immunoglobulin-like fold and provides new insights into pilus assembly, function and adhesion.
- Department of Ultrastructure, Institute for Molecular Biology, Vrije Universiteit Brussel, Vlaams Interuniversitair Instituut voor Biotechnologie, Brussels, Belgium.
Organizational Affiliation: