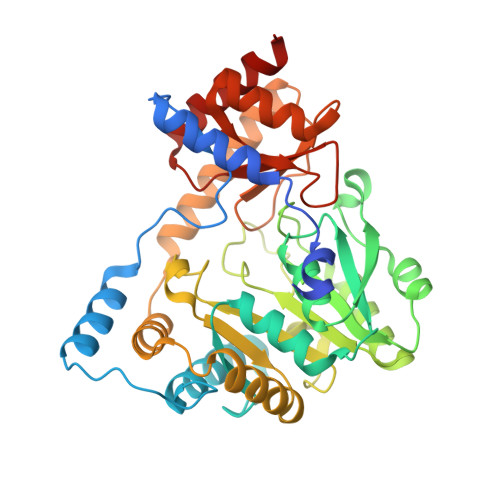
Crystal structure of an aspartate aminotransferase (TM1255) from Thermotoga maritima at 1.90 A resolution
Schwarzenbacher, R., Jaroszewski, L., von Delft, F., Abdubek, P., Ambing, E., Biorac, T., Brinen, L.S., Canaves, J.M., Cambell, J., Chiu, H.J., Dai, X., Deacon, A.M., DiDonato, M., Elsliger, M.A., Eshagi, S., Floyd, R., Godzik, A., Grittini, C., Grzechnik, S.K., Hampton, E., Karlak, C., Klock, H.E., Koesema, E., Kovarik, J.S., Kreusch, A., Kuhn, P., Lesley, S.A., Levin, I., McMullan, D., McPhillips, T.M., Miller, M.D., Morse, A., Moy, K., Ouyang, J., Page, R., Quijano, K., Robb, A., Spraggon, G., Stevens, R.C., van den Bedem, H., Velasquez, J., Vincent, J., Wang, X., West, B., Wolf, G., Xu, Q., Hodgson, K.O., Wooley, J., Wilson, I.A.(2004) Proteins 55: 759-763
- PubMed: 15103638 Search on PubMed
- DOI: https://doi.org/10.1002/prot.10646
- Primary Citation Related Structures:
1O4S - Joint Center for Structural Genomics, La Jolla, CA 92037, USA.
Organizational Affiliation: