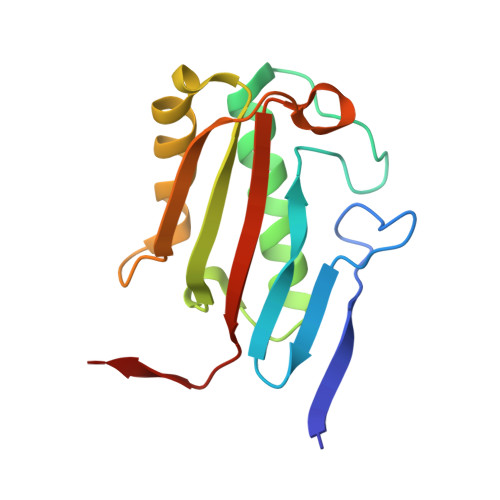
Structure and oligomeric state of the mammalian tumour-associated antigen UK114.
Deriu, D., Briand, C., Mistiniene, E., Naktinis, V., Grutter, M.G.(2003) Acta Crystallogr D Biol Crystallogr 59: 1676-1678
- PubMed: 12925811 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444903014306
- Primary Citation Related Structures:
1NQ3 - PubMed Abstract:
The tumour-associated antigen UK114, isolated from goat liver, belongs to the YER057c/YIL051c/YjgF protein family, which has members in both the prokaryotes and eukaryotes. The crystal structure of a mammalian representative, goat UK114, was determined, revealing a trimeric arrangement in the crystal. It was confirmed by ultracentrifugation that UK114 is a trimer in solution. These results are in agreement with the published structures of homologues from unicellular organisms, but contrast with those reported for the rat homologue of UK114, for which a dimeric quaternary structure was proposed.
- Institute of Biochemistry, University of Zurich, Winterthurerstrasse 190, CH-8057 Zurich, Switzerland.
Organizational Affiliation: