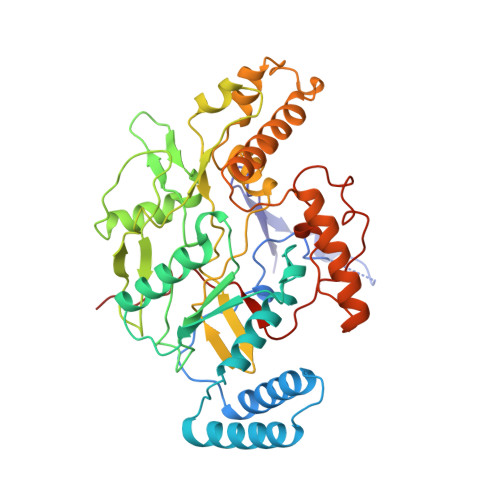
Structure of nitric oxide synthase oxygenase dimer with pterin and substrate.
Crane, B.R., Arvai, A.S., Ghosh, D.K., Wu, C., Getzoff, E.D., Stuehr, D.J., Tainer, J.A.(1998) Science 279: 2121-2126
- PubMed: 9516116 Search on PubMed
- DOI: https://doi.org/10.1126/science.279.5359.2121
- Primary Citation Related Structures:
1NOD, 2NOD, 3NOD - PubMed Abstract:
Crystal structures of the murine cytokine-inducible nitric oxide synthase oxygenase dimer with active-center water molecules, the substrate L-arginine (L-Arg), or product analog thiocitrulline reveal how dimerization, cofactor tetrahydrobiopterin, and L-Arg binding complete the catalytic center for synthesis of the essential biological signal and cytotoxin nitric oxide. Pterin binding refolds the central interface region, recruits new structural elements, creates a 30 angstrom deep active-center channel, and causes a 35 degrees helical tilt to expose a heme edge and the adjacent residue tryptophan-366 for likely reductase domain interactions and caveolin inhibition. Heme propionate interactions with pterin and L-Arg suggest that pterin has electronic influences on heme-bound oxygen. L-Arginine binds to glutamic acid-371 and stacks with heme in an otherwise hydrophobic pocket to aid activation of heme-bound oxygen by direct proton donation and thereby differentiate the two chemical steps of nitric oxide synthesis.
- Department of Molecular Biology and Skaggs Institute for Chemical Biology, The Scripps Research Institute, La Jolla, CA 92037, USA.
Organizational Affiliation: