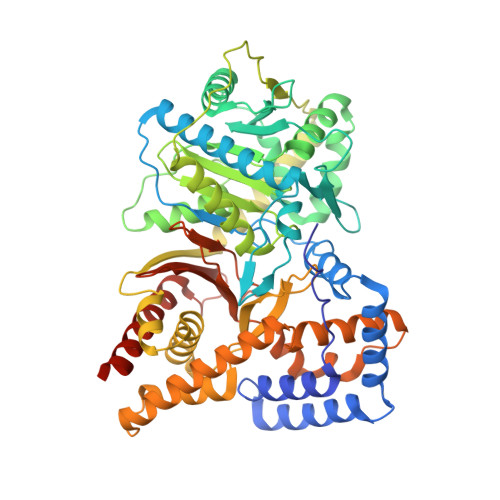
Structure of Human Carnitine Acetyltransferase. Molecular Basis for Fatty Acyl Transfer
Wu, D., Govindasamy, L., Lian, W., Gu, Y., Kukar, T., Agbandje-McKenna, M., McKenna, R.(2003) J Biological Chem 278: 13159-13165
- PubMed: 12562770 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M212356200
- Primary Citation Related Structures:
1NM8 - PubMed Abstract:
Carnitine acyltransferases are a family of ubiquitous enzymes that play a pivotal role in cellular energy metabolism. We report here the x-ray structure of human carnitine acetyltransferase to a 1.6-A resolution. This structure reveals a monomeric protein of two equally sized alpha/beta domains. Each domain is shown to have a partially similar fold to other known but oligomeric enzymes that are also involved in group-transfer reactions. The unique monomeric arrangement of the two domains constitutes a central narrow active site tunnel, indicating a likely universal feature for all members of the carnitine acyltransferase family. Superimposition of the substrate complex of a related protein, dihydrolipoyl trans-acetylase, reveals that both substrates localize to the active site tunnel of human carnitine acetyltransferase, suggesting the location of the ligand binding sites for carnitine and coenzyme A. Most significantly, this structure provides critical insights into the molecular basis for fatty acyl chain transfer and a possible common mechanism among a wide range of acyltransferases utilizing a catalytic dyad.
- Department of Medicinal Chemistry, College of Pharmacy, University of Florida, Gainesville, Florida 32610, USA.
Organizational Affiliation: