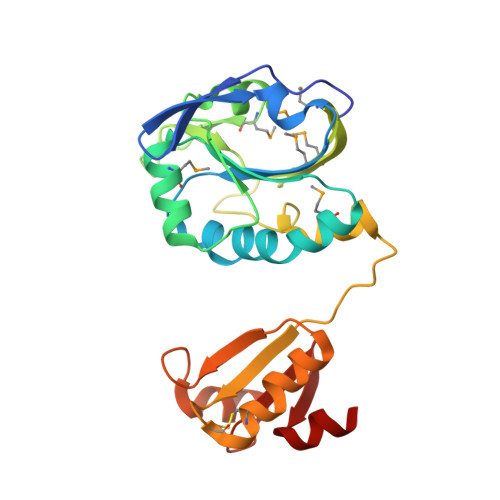
The Tetrameric Structure of Haemophilus influenza Hybrid Prx5 Reveals Interactions between Electron Donor and Acceptor Proteins.
Kim, S.J., Woo, J.R., Hwang, Y.S., Jeong, D.G., Shin, D.H., Kim, K., Ryu, S.E.(2003) J Biological Chem 278: 10790-10798
- PubMed: 12529327 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M209553200
- Primary Citation Related Structures:
1NM3 - PubMed Abstract:
Cellular redox control is often mediated by oxidation and reduction of cysteine residues in the redox-sensitive proteins, where thioredoxin and glutaredoxin (Grx) play as electron donors for the oxidized proteins. Despite the importance of protein-protein interactions between the electron donor and acceptor proteins, there has been no structural information for the interaction of thioredoxin or Grx with natural target proteins. Here, we present the crystal structure of a novel Haemophilus influenza peroxiredoxin (Prx) hybrid Prx5 determined at 2.8-A resolution. The structure reveals that hybrid Prx5 forms a tightly associated tetramer where active sites of Prx and Grx domains of different monomers interact with each other. The Prx-Grx interface comprises specific charge interactions surrounded by weak interactions, providing insight into the target recognition mechanism of Grx. The tetrameric structure also exhibits a flexible active site and alternative Prx-Grx interactions, which appear to facilitate the electron transfer from Grx to Prx domain. Differences of electron donor binding surfaces in Prx proteins revealed by an analysis based on the structural information explain the electron donor specificities of various Prx proteins.
- Center for Cellular Switch Protein Structure, Korea Research Institute of Bioscience and Biotechnology, 52 Euh-eun-dong, Yusong-gu, Daejon 305-806, South Korea.
Organizational Affiliation: