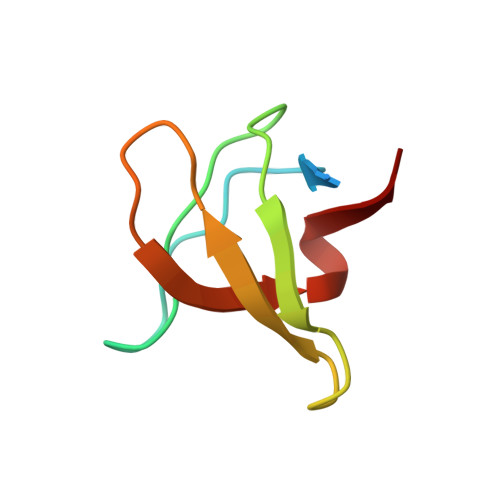
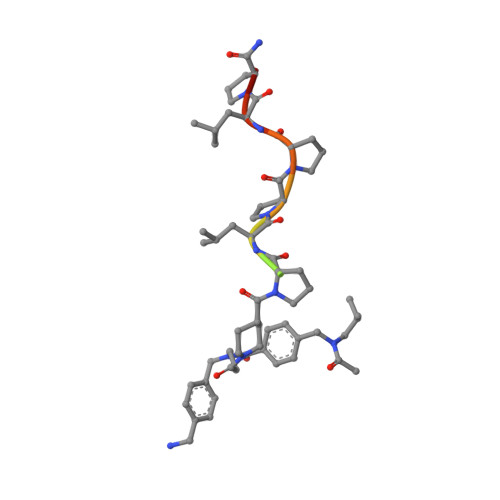
Molecular basis for the binding of SH3 ligands with non-peptide elements identified by combinatorial synthesis.
Feng, S., Kapoor, T.M., Shirai, F., Combs, A.P., Schreiber, S.L.(1996) Chem Biol 3: 661-670
- PubMed: 8807900 Search on PubMed
- DOI: https://doi.org/10.1016/s1074-5521(96)90134-9
- Primary Citation Related Structures:
1NLO, 1NLP - PubMed Abstract:
Protein-structure-based combinatorial chemistry has recently been used to discover several ligands containing non-peptide binding elements to the Src SH3 domain. The encoded library used has the form Cap-M1-M2-M3-PLPPLP, in which the Cap and Mi's are composed of a diverse set of organic monomers. The PLPPLP portion provided a structural bias directing the non-peptide fragment Cap-M1-M2-M3 to the SH3 specificity pocket. Fifteen ligands were selected from > 1.1 million distinct compounds. The structural basis for selection was unknown. The solution structures of the Src SH3 domain complexed with two ligands containing non-peptide elements selected from the library were determined by multidimensional NMR spectroscopy. The non-peptide moieties of the ligands interact with the specificity pocket of Src SH3 domain differently from peptides complexed with SH3 domains. Structural information about the ligands was used to design various homologs, whose affinities for the SH3 domain were measured. The results provide a structural basis for understanding the selection of a few optimal ligands from a large library. The cycle of protein-structure-based combinatorial chemistry followed by structure determination of the few highest affinity ligands provides a powerful new tool for the field of molecular recognition.
- Howard Hughes Medical Institute, Department of Chemistry and Chemical Biology, Harvard University, MA 02138, USA.
Organizational Affiliation: