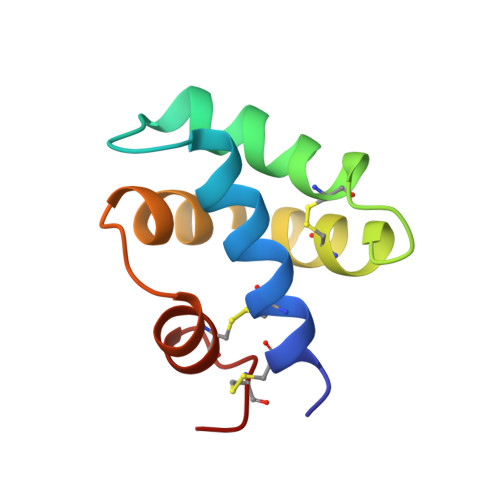
Saposin fold revealed by the NMR structure of NK-lysin.
Liepinsh, E., Andersson, M., Ruysschaert, J.M., Otting, G.(1997) Nat Struct Biol 4: 793-795
- PubMed: 9334742 Search on PubMed
- DOI: https://doi.org/10.1038/nsb1097-793
- Primary Citation Related Structures:
1NKL - PubMed Abstract:
NK-lysin is the first representative of a family of sequence related proteins--saposins, surfactant-associated protein B, pore forming amoeba proteins, and domains of acid sphingomyelinase, acyloxyacylhydrolase and plant aspartic proteinases--for which a structure has been determined.