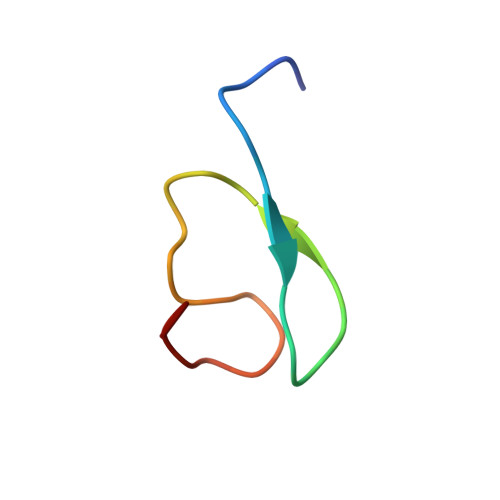
Structure and ubiquitin interactions of the conserved zinc finger domain of Npl4.
Wang, B., Alam, S.L., Meyer, H.H., Payne, M., Stemmler, T.L., Davis, D.R., Sundquist, W.I.(2003) J Biological Chem 278: 20225-20234
- PubMed: 12644454 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M300459200
- Primary Citation Related Structures:
1NJ3 - PubMed Abstract:
Ubiquitylated proteins are directed into a large number of different cellular pathways through interactions with effector proteins that contain conserved ubiquitin binding motifs. Here, we report the solution structure and ubiquitin binding properties of one such motif, the Npl4 zinc finger or RanBP2/Nup358 zinc finger (NZF) domain. Npl4 NZF forms a compact module composed of four antiparallel beta-strands linked by three ordered loops. A single zinc ion is coordinated by four conserved cysteines from the first and third loops, which form two rubredoxin knuckles. Npl4 NZF binds specifically, but weakly, to free ubiquitin using a conserved 13TF14 dipeptide to interact with the "Ile-44" surface of ubiquitin. Our studies reveal the structure of this versatile class of protein binding domains and provide a means for identifying the subset of NZF domains likely to bind ubiquitin.
- Department of Biochemistry, University of Utah, Salt Lake City 84132, USA.
Organizational Affiliation: