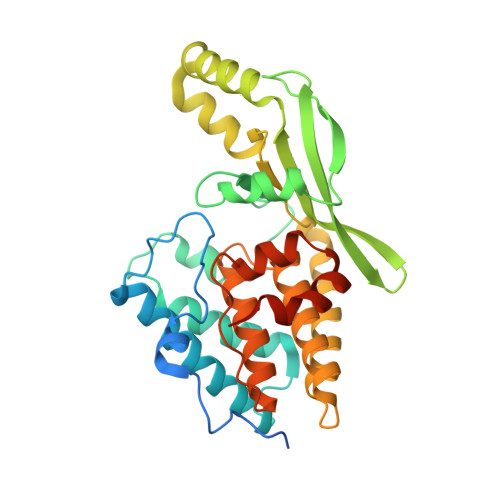
Crystal Structure of the Type III Effector AvrB from Pseudomonas syringae.
Lee, C.C., Wood, M.D., Ng, K., Luginbuhl, P., Spraggon, G., Katagiri, F.(2004) Structure 12: 487-494
- PubMed: 15016364 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2004.02.013
- Primary Citation Related Structures:
1NH1 - PubMed Abstract:
AvrB is a Pseudomonas syringae type III effector protein that is translocated into host plant cells during attempted pathogenesis. Arabidopsis harboring the corresponding resistance protein RPM1 can detect AvrB and mount a rapid host defense response, thus avoiding active infection. In the plant cell, AvrB induces phosphorylation of RIN4, a key component in AvrB/RPM1 recognition. Although the AvrB/RPM1 system is among the best characterized of the numerous bacterial effector/plant resistance protein systems involved in plant disease resistance and pathogenesis, the details of the molecular recognition mechanism are still unclear. To gain further insights, the crystal structure of AvrB was determined. The 2.2 A structure exhibits a novel mixed alpha/beta bilobal fold. Aided by the structural information, we demonstrate that one lobe is the determinant of AvrB/RPM1 recognition specificity. This structural information and preliminary structure-function studies provide a framework for the future understanding of AvrB function on the molecular level.
- Genomics Institute of the Novartis Research Foundation, 10675 John Jay Hopkins Drive, San Diego, CA 92121, USA.
Organizational Affiliation: