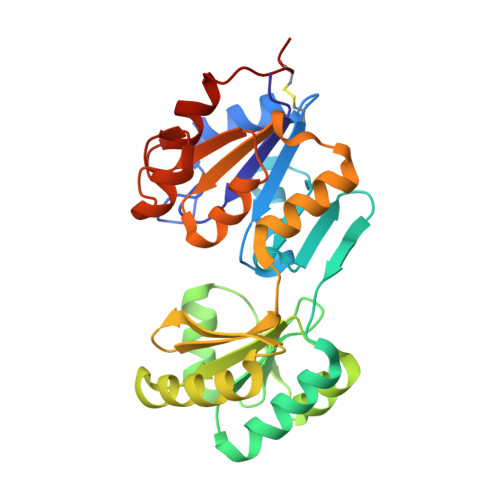
Crystal structure of a phosphatase with a unique substrate binding domain from Thermotoga maritima
Shin, D.H., Roberts, A., Jancarik, J., Yokota, H., Kim, R., Wemmer, D.E., Kim, S.H.(2003) Protein Sci 12: 1464-1472
- PubMed: 12824492 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.0302703
- Primary Citation Related Structures:
1NF2 - PubMed Abstract:
We have determined the crystal structure of a phosphatase with a unique substrate binding domain from Thermotoga maritima, TM0651 (gi 4981173), at 2.2 A resolution by selenomethionine single-wavelength anomalous diffraction (SAD) techniques. TM0651 is a member of the haloacid dehalogenase (HAD) superfamily, with sequence homology to trehalose-6-phosphate phosphatase and sucrose-6(F)-phosphate phosphohydrolase. Selenomethionine labeled TM0651 crystallized in space group C2 with three monomers per asymmetric unit. Each monomer has approximate dimensions of 65 x 40 x 35 A(3), and contains two domains: a domain of known hydrolase fold characteristic of the HAD family, and a domain with a new tertiary fold consisting of a six-stranded beta-sheet surrounded by four alpha-helices. There is one disulfide bond between residues Cys35 and Cys265 in each monomer. One magnesium ion and one sulfate ion are bound in the active site. The superposition of active site residues with other HAD family members indicates that TM0651 is very likely a phosphatase that acts through the formation of a phosphoaspartate intermediate, which is supported by both NMR titration data and a biochemical assay. Structural and functional database searches and the presence of many aromatic residues in the interface of the two domains suggest the substrate of TM0651 is a carbohydrate molecule. From the crystal structure and NMR data, the protein likely undergoes a conformational change upon substrate binding.
- Physical Biosciences Division, Lawrence Berkeley National Laboratory, Berkeley, CA 94720, USA.
Organizational Affiliation: