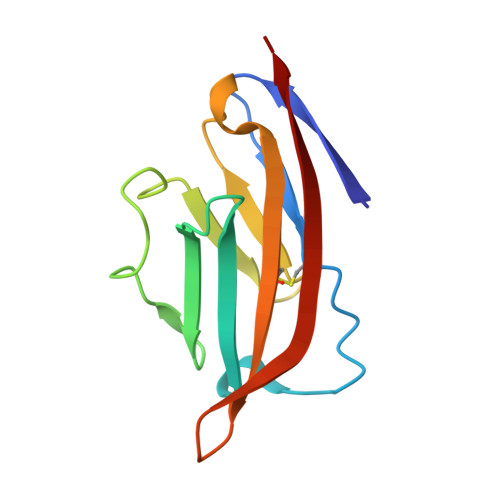
Crystal Structure of the Receptor-Binding Domain of Human B7-2: Insights into Organization and Signaling
Zhang, X., Schwartz, J.D., Almo, S.C., Nathenson, S.G.(2003) Proc Natl Acad Sci U S A 100: 2586-2591
- PubMed: 12606712 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.252771499
- Primary Citation Related Structures:
1NCN - PubMed Abstract:
B7-1 and B7-2 are homologous costimulatory ligands expressed on the surfaces of antigen-presenting cells. Their interactions with CD28/CTLA-4 receptors expressed on T cell surfaces are crucial for the proper regulation of T cell activity. B7-1 and B7-2 display distinct roles in immune regulation, although they are usually considered to have redundant functions. Here, we report the crystal structure of the receptor-binding (Ig V-type) domain of human B7-2 at 2.7-A resolution. Structures of unliganded and liganded B7-1 and B7-2 suggest a physical-chemical basis for the observed functional similarities and differences between these two costimulatory ligands. Of particular note, whereas the majority of the residues mediating B7-1 dimerization are hydrophobic, the B7-2 dimer observed in the B7-2/CTLA-4 complex displays a very hydrophilic dimer interface. These differences provide a mechanism for preventing the formation of B7-1/B7-2 heterodimers. The divergence at the putative dimer interface is also consistent with the lower tendency of B7-2 to dimerize, as shown by the monomeric state of unliganded B7-2 both in solution and crystalline form, and may result in detailed differences in signaling mechanisms associated with B7-1 and B7-2.
- Department of Cell Biology, Albert Einstein College of Medicine, 1300 Morris Park Avenue, Bronx, NY 10461, USA.
Organizational Affiliation: