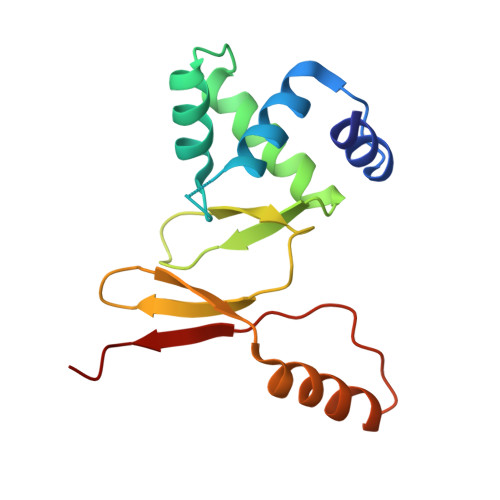
Architecture of a protein central to iron homeostatis: Crystal structure and spectroscopic analysis of the Ferric uptake regulator
Pohl, E., Haller, J.C., Mijovilovich, A., Meyer-Klaucke, W., Garman, E., Vasil, M.L.(2003) Mol Microbiol 47: 903-915
- PubMed: 12581348 Search on PubMed
- DOI: https://doi.org/10.1046/j.1365-2958.2003.03337.x
- Primary Citation Related Structures:
1MZB - PubMed Abstract:
Iron is an essential element for almost all organisms, although an overload of this element results in toxicity because of the formation of hydroxyl radicals. Consequently, most living entities have developed sophisticated mechanisms to control their intracellular iron concentration. In many bacteria, including the opportunistic pathogen Pseudomonas aeruginosa, this task is performed by the ferric uptake regulator (Fur). Fur controls a wide variety of basic physiological processes including iron uptake systems and the expression of exotoxin A. Here, we present the first crystal structure of Fur from P. aeruginosa in complex with Zn2+ determined at a resolution of 1.8 A. Furthermore, X-ray absorption spectroscopic measurements and microPIXE analysis were performed in order to characterize the distinct zinc and iron binding sites in solution. The combination of these complementary techniques enables us to present a model for the activation and DNA binding of the Fur protein.
- European Molecular Biology Laboratory, Hamburg Outstation, Notkestr. 85, D-22603 Hamburg, Germany.
Organizational Affiliation: