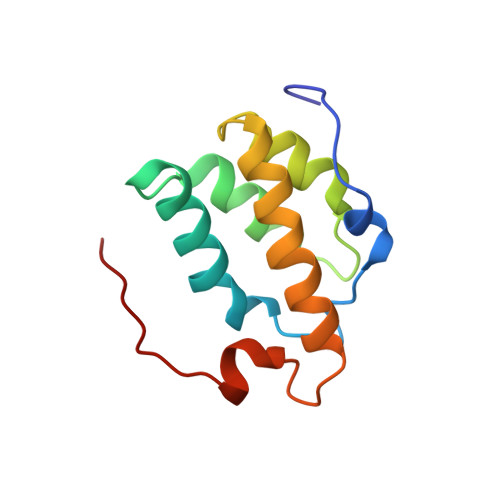
Structure and function of the PWI motif: a novel nucleic acid-binding domain that facilitates pre-mRNA processing.
Szymczyna, B.R., Bowman, J., McCracken, S., Pineda-Lucena, A., Lu, Y., Cox, B., Lambermon, M., Graveley, B.R., Arrowsmith, C.H., Blencowe, B.J.(2003) Genes Dev 17: 461-475
- PubMed: 12600940 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1101/gad.1060403
- Primary Citation Related Structures:
1MP1 - PubMed Abstract:
The PWI motif is a highly conserved domain of unknown function in the SRm160 splicing and 3'-end cleavage-stimulatory factor, as well as in several other known or putative pre-mRNA processing components. We show here that the PWI motif is a new type of RNA/DNA-binding domain that has an equal preference for single- and double-stranded nucleic acids. Deletion of the motif prevents SRm160 from binding RNA and stimulating 3'-end cleavage, and its substitution with a heterologous RNA-binding domain restores these functions. The NMR solution structure of the SRm160-PWI motif reveals a novel, four-helix bundle and represents the first example of an alpha-helical fold that can bind single-stranded (ss)RNA. Structure-guided mutagenesis indicates that the same surface is involved in RNA and DNA binding and requires the cooperative action of a highly conserved, adjacent basic region. Thus, the PWI motif is a novel type of nucleic acid-binding domain that likely has multiple important functions in pre-mRNA processing, including SRm160-dependent stimulation of 3'-end formation.
- Ontario Cancer Institute, University of Toronto, Toronto, Ontario, Canada M5G 2M9.
Organizational Affiliation: