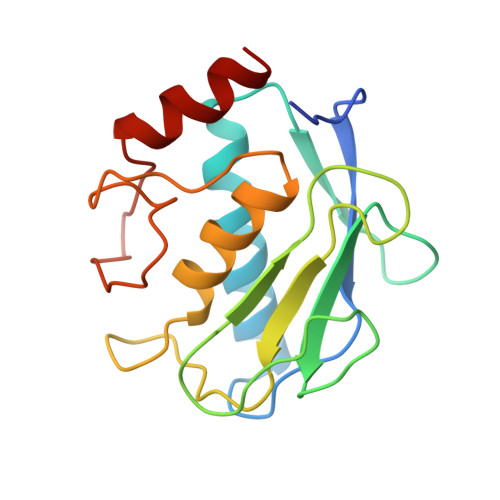
Structure determination and analysis of human neutrophil collagenase complexed with a hydroxamate inhibitor.
Grams, F., Crimmin, M., Hinnes, L., Huxley, P., Pieper, M., Tschesche, H., Bode, W.(1995) Biochemistry 34: 14012-14020
- PubMed: 7577999 Search on PubMed
- DOI: https://doi.org/10.1021/bi00043a007
- Primary Citation Related Structures:
1MMB - PubMed Abstract:
Matrix metalloproteinases are a family of zinc endopeptidases involved in tissue remodeling. They have been implicated in various disease processes including metastasis, joint destruction, and neurodegeneration. Human neutrophil collagenase (HNC, MMP-8) represents one of the three "interstitial" collagenases that cleave triple-helical collagens types I, II, and III. Its 163-residue catalytic domain (Met80 to Gly242) has been expressed in Escherichia coli and crystallized as a noncovalent complex with the hydroxamate inhibitor batimastat. The crystal structure, refined to 2.1 A, demonstrates that batimastat binds to the S1-S2' sites and coordinates to the catalytic zinc in a bidentate manner via the hydroxyl and carbonyl oxygens of the hydroxamate group. The batimastat-collagenase complex is described in detail, and the activities of batimastat analogues are discussed in the light of the protein-inhibitor interactions revealed by the crystallography studies.
- Abteilung für Strukturforschung, Max-Planck-Institut für Biochemie, Martinsried, Germany.
Organizational Affiliation: