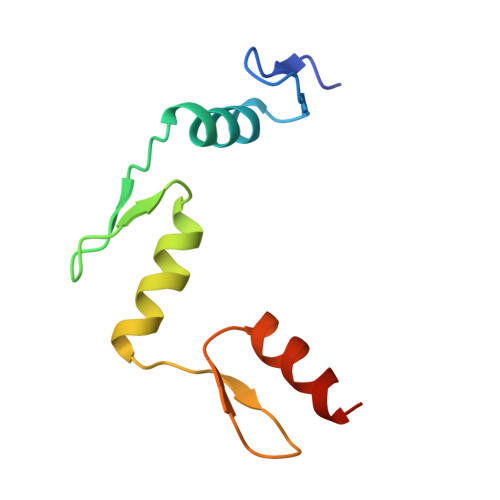
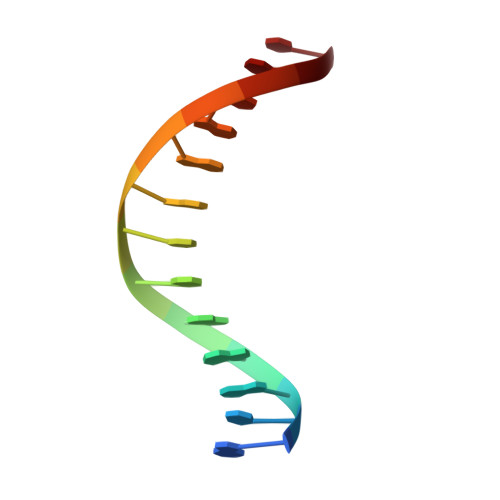
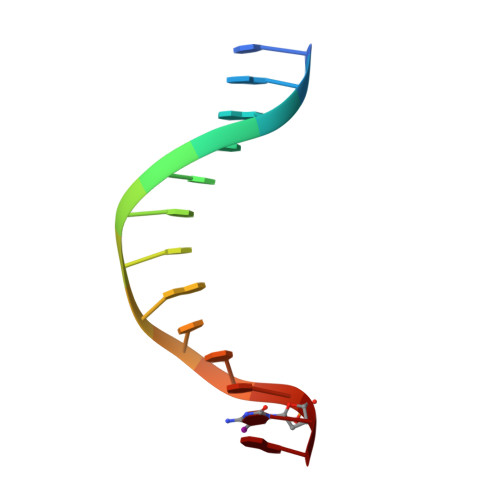
A 2.2 A Resolution Crystal Structure of a Designed Zinc Finger Protein Bound to DNA
Kim, C.A., Berg, J.M.(1996) Nat Struct Biol 3: 940-945
- PubMed: 8901872 Search on PubMed
- DOI: https://doi.org/10.1038/nsb1196-940
- Primary Citation Related Structures:
1MEY - PubMed Abstract:
Considerable recent effort has been devoted to the design and selection of sequence-specific DNA binding proteins based on tandem arrays of Cys2His2 zinc finger domains. While the DNA binding properties of these designed proteins have been studied extensively, the structural basis for site-specific binding has not been examined experimentally. Here we report the crystal structure of a complex between a protein comprised of three consensus-sequence-based zinc finger domains and an oligonucleotide corresponding to a favourable DNA binding site. This structure reveals relatively simple modular interactions and structural adaptations that compensate for differences in contact residue side-chain lengths.
- Department of Chemistry, Johns Hopkins University, Baltimore, Maryland 21218, USA.
Organizational Affiliation: