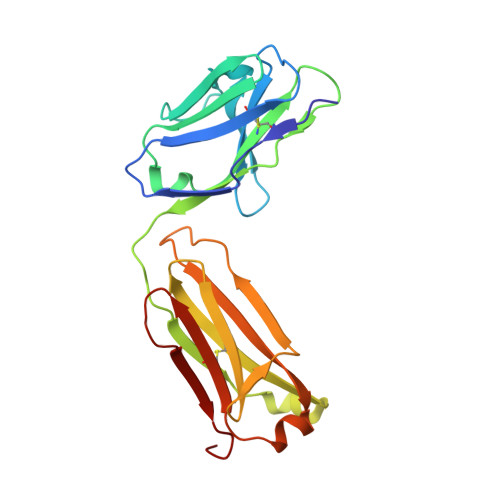
Crystal structure to 2.45 A resolution of a monoclonal Fab specific for the Brucella A cell wall polysaccharide antigen.
Rose, D.R., Przybylska, M., To, R.J., Kayden, C.S., Oomen, R.P., Vorberg, E., Young, N.M., Bundle, D.R.(1993) Protein Sci 2: 1106-1113
- PubMed: 8358294 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.5560020705
- Primary Citation Related Structures:
1MAM - PubMed Abstract:
The atomic structure of an antibody antigen-binding fragment (Fab) at 2.45 A resolution shows that polysaccharide antigen conformation and Fab structure dictated by combinatorial diversity and domain association are responsible for the fine specificity of the Brucella-specific antibody, YsT9.1. It discriminates the Brucella abortus A antigen from the nearly identical Brucella melitensis M antigen by forming a groove-type binding site, lined with tyrosine residues, that accommodates the rodlike A antigen but excludes the kinked structure of the M antigen, as envisioned by a model of the antigen built into the combining site. The variable-heavy (VH) and variable-light (VL) domains are derived from genes closely related to two used in previously solved structures, M603 and R19.9, respectively. These genes combine in YsT9.1 to form an antibody of totally different specificity. Comparison of this X-ray structure with a previously built model of the YsT9.1 combining site based on these homologies highlights the importance of VL:VH association as a determinant of specificity and suggests that small changes at the VL:VH interface, unanticipated in modeling, may cause significant modulation of binding-site properties.
- Ontario Cancer Institute, University of Toronto, Canada.
Organizational Affiliation: