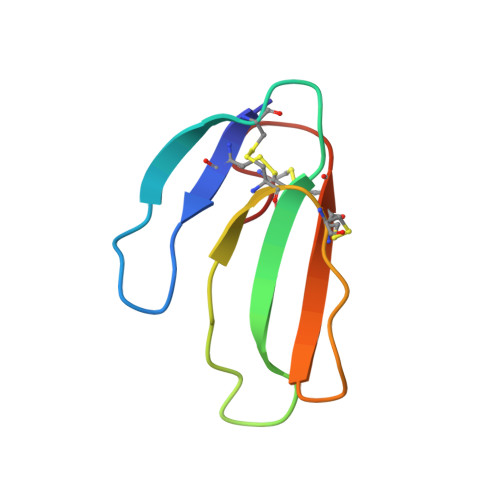
Acetylcholinesterase inhibition by fasciculin: crystal structure of the complex.
Bourne, Y., Taylor, P., Marchot, P.(1995) Cell 83: 503-512
- PubMed: 8521480 Search on PubMed
- DOI: https://doi.org/10.1016/0092-8674(95)90128-0
- Primary Citation Related Structures:
1MAH - PubMed Abstract:
The crystal structure of the snake toxin fasciculin, bound to mouse acetylcholinesterase (mAChE), at 3.2 A resolution reveals a synergistic three-point anchorage consistent with the picomolar dissociation constant of the complex. Loop II of fasciculin contains a cluster of hydrophobic residues that interact with the peripheral anionic site of the enzyme and sterically occlude substrate access to the catalytic site. Loop I fits in a crevice near the lip of the gorge to maximize the surface area of contact of loop II at the gorge entry. The fasciculin core surrounds a protruding loop on the enzyme surface and stabilizes the whole assembly. Upon binding of fasciculin, subtle structural rearrangements of AChE occur that could explain the observed residual catalytic activity of the fasciculin-enzyme complex.
- Department of Pharmacology, University of California, San Diego, La Jolla 92093-0636, USA.
Organizational Affiliation: