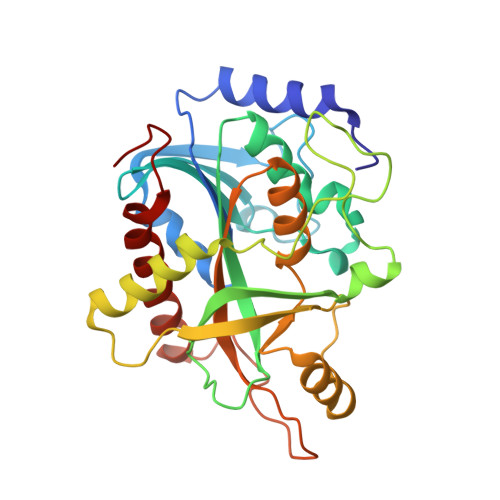
Crystal structure of human purine nucleoside phosphorylase at 2.3A resolution.
de Azevedo, W.F., Canduri, F., dos Santos, D.M., Silva, R.G., de Oliveira, J.S., de Carvalho, L.P., Basso, L.A., Mendes, M.A., Palma, M.S., Santos, D.S.(2003) Biochem Biophys Res Commun 308: 545-552
- PubMed: 12914785 Search on PubMed
- DOI: https://doi.org/10.1016/s0006-291x(03)01431-1
- Primary Citation Related Structures:
1M73 - PubMed Abstract:
Purine nucleoside phosphorylase (PNP) catalyzes the phosphorolysis of the N-ribosidic bonds of purine nucleosides and deoxynucleosides. In human, PNP is the only route for degradation of deoxyguanosine and genetic deficiency of this enzyme leads to profound T-cell mediated immunosuppression. PNP is therefore a target for inhibitor development aiming at T-cell immune response modulation and its low resolution structure has been used for drug design. Here we report the structure of human PNP solved to 2.3A resolution using synchrotron radiation and cryocrystallographic techniques. This structure allowed a more precise analysis of the active site, generating a more reliable model for substrate binding. The higher resolution data allowed the identification of water molecules in the active site, which suggests binding partners for potential ligands. Furthermore, the present structure may be used in the new structure-based design of PNP inhibitors.
- Departamento de Física, UNESP, São José do Rio Preto, SP, Brazil. walterfa@df.ibilce.unesp.br
Organizational Affiliation: