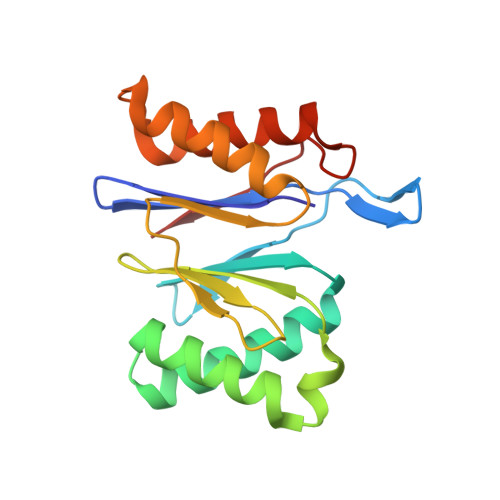
Isolation and characterization of the prokaryotic proteasome homolog HslVU (ClpQY) from Thermotoga maritima and the crystal structure of HslV.
Song, H.K., Bochtler, M., Azim, M.K., Hartmann, C., Huber, R., Ramachandran, R.(2003) Biophys Chem 100: 437-452
- PubMed: 12646382 Search on PubMed
- DOI: https://doi.org/10.1016/s0301-4622(02)00297-1
- Primary Citation Related Structures:
1M4Y - PubMed Abstract:
Heat-shock locus VU (HslVU) is an ATP-dependent proteolytic system and a prokaryotic homolog of the proteasome. It consists of HslV, the protease, and HslU, the ATPase and chaperone. We have cloned, sequenced and expressed both protein components from the hyperthermophile Thermotoga maritima. T. maritima HslU hydrolyzes a variety of nucleotides in a temperature-dependent manner, with the optimum lying between 75 and 80 degrees C. It is also nucleotide-unspecific for activation of HslV against amidolytic and caseinolytic activity. The Escherichia coli and T. maritima HslU proteins mutually stimulate HslV proteins from both sources, suggesting a conserved activation mechanism. The crystal structure of T. maritima HslV was determined and refined to 2.1-A resolution. The structure of the dodecameric enzyme is well conserved compared to those from E. coli and Haemophilus influenzae. A comparison of known HslV structures confirms the presence of a cation-binding site, although its exact role in the proteolytic mechanism of HslV remains unclear. Amongst factors responsible for the thermostability of T. maritima HslV, extensive ionic interactions/salt-bridge networks, which occur specifically in the T. maritima enzyme in comparison to its mesophilic counterparts, seem to play an important role.
- Abteilung Strukturforschung, Max-Planck-Institut für Biochemie, Am Klopferspitz 18a, D-82152 Planegg-Martinsried, Germany.
Organizational Affiliation: