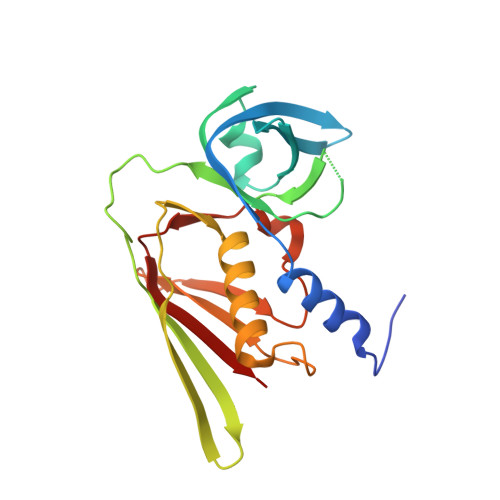
The three-dimensional structure of a superantigen-like protein, SET3, from a pathogenicity island of the Staphylococcus aureus genome
Arcus, V.L., Langley, R., Proft, T., Fraser, J.D., Baker, E.N.(2002) J Biological Chem 277: 32274-32281
- PubMed: 12082105 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M203914200
- Primary Citation Related Structures:
1M4V - PubMed Abstract:
The staphylococcal enterotoxin-like toxins (SETs) are a family of proteins encoded within the Staphylococcus aureus genome that were identified by their similarity to the well described bacterial superantigens. The first crystal structure of a member of the SET family, SET3, has been determined to 1.9 A (R = 0.205, R(free) = 0.240) and reveals a fold characteristic of the superantigen family but with significant differences. The SET proteins are secreted at varying levels by staphylococcal isolates, and seroconversion studies of normal individuals indicate that they are strongly antigenic to humans. Recombinant SETs do not exhibit any of the properties expected of superantigens such as major histocompatibility complex class II binding or broad T-cell activation, suggesting they have an entirely different function. The fact that the whole gene family is clustered within the pathogenicity island SaIn2 of the S. aureus genome suggests that they are involved in host/pathogen interactions.
- School of Biological Sciences, University of Auckland, Private Bag 92-019 Auckland, New Zealand.
Organizational Affiliation: