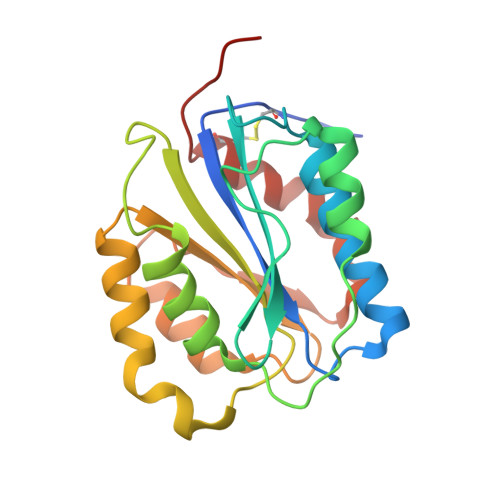
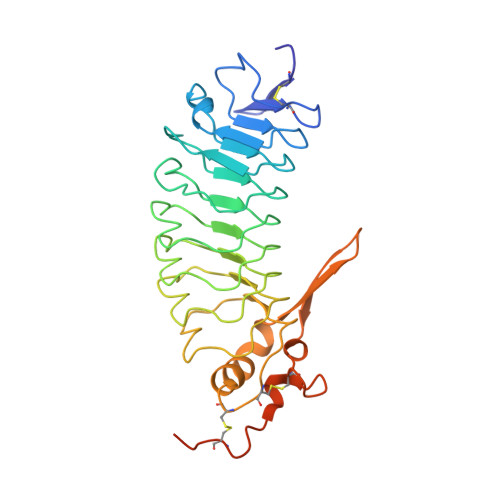
Structures of glycoprotein Ibalpha and its complex with von Willebrand factor A1 domain.
Huizinga, E.G., Tsuji, S., Romijn, R.A., Schiphorst, M.E., de Groot, P.G., Sixma, J.J., Gros, P.(2002) Science 297: 1176-1179
- PubMed: 12183630 Search on PubMed
- DOI: https://doi.org/10.1126/science.107355
- Primary Citation Related Structures:
1M0Z, 1M10 - PubMed Abstract:
Transient interactions of platelet-receptor glycoprotein Ibalpha (GpIbalpha) and the plasma protein von Willebrand factor (VWF) reduce platelet velocity at sites of vascular damage and play a role in haemostasis and thrombosis. Here we present structures of the GpIbalpha amino-terminal domain and its complex with the VWF domain A1. In the complex, GpIbalpha wraps around one side of A1, providing two contact areas bridged by an area of solvated charge interaction. The structures explain the effects of gain-of-function mutations related to bleeding disorders and provide a model for shear-induced activation. These detailed insights into the initial interactions in platelet adhesion are relevant to the development of antithrombotic drugs.
- Department of Crystal and Structural Chemistry, Bijvoet Center for Biomolecular Research, Utrecht University, Padualaan 8, 3584 CH Utrecht, Netherlands. e.g.huizinga@chem.uu.nl
Organizational Affiliation: