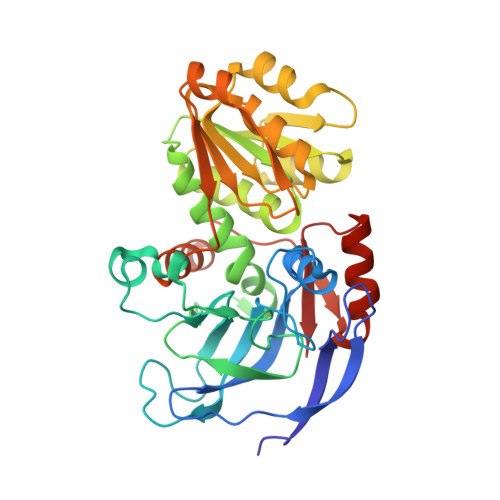
The ternary complex of Pseudomonas aeruginosa alcohol dehydrogenase with NADH and ethylene glycol.
Levin, I., Meiri, G., Peretz, M., Burstein, Y., Frolow, F.(2004) Protein Sci 13: 1547-1556
- PubMed: 15152088 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.03531404
- Primary Citation Related Structures:
1LLU - PubMed Abstract:
Pseudomonas aeruginosa alcohol dehydrogenase (PaADH; ADH, EC 1.1.1.1) catalyzes the reversible oxidation of primary and secondary alcohols to the corresponding aldehydes and ketones, using NAD as coenzyme. We crystallized the ternary complex of PaADH with its coenzyme and a substrate molecule and determined its structure at a resolution of 2.3 A, using the molecular replacement method. The PaADH tetramer comprises four identical chains of 342 amino acid residues each and obeys ~222-point symmetry. The PaADH monomer is structurally similar to alcohol dehydrogenase monomers from vertebrates, archaea, and bacteria. The stabilization of the ternary complex of PaADH, the coenzyme, and the poor substrate ethylene glycol (k(cat) = 4.5 sec(-1); Km > 200 mM) was due to the blocked exit of the coenzyme in the crystalline state, combined with a high (2.5 M) concentration of the substrate. The structure of the ternary complex presents the precise geometry of the Zn coordination complex, the proton-shuttling system, and the hydride transfer path. The ternary complex structure also suggests that the low efficiency of ethylene glycol as a substrate results from the presence of a second hydroxyl group in this molecule.
- Department of Organic Chemistry, Weizmann Institute of Science, 76100 Rehovot, Israel.
Organizational Affiliation: