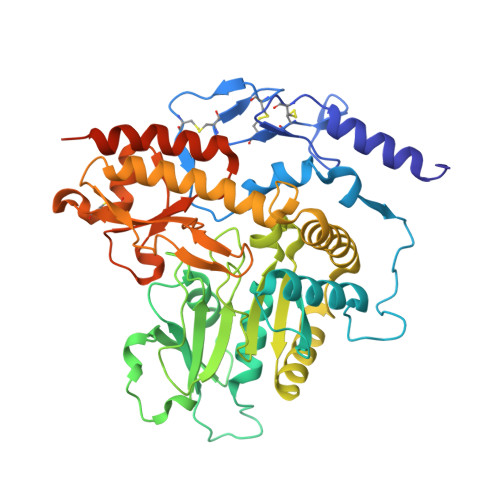
The active principle of garlic at atomic resolution
Kuettner, E.B., Hilgenfeld, R., Weiss, M.S.(2002) J Biological Chem 277: 46402-46407
- PubMed: 12235163 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M208669200
- Primary Citation Related Structures:
1LK9 - PubMed Abstract:
Despite the fact that many cultures around the world value and utilize garlic as a fundamental component of their cuisine as well as of their medicine cabinets, relatively little is known about the plant's protein configuration that is responsible for the specific properties of garlic. Here, we report the three-dimensional structure of the garlic enzyme alliinase at 1.5 A resolution. Alliinase constitutes the major protein component in garlic bulbs, and it is able to cleave carbon-sulfur bonds. The active enzyme is a pyridoxal-5'-phosphate-dependent homodimeric glycoprotein and belongs to the class I family of pyridoxal-5'-phosphate-dependent enzymes. In addition, it contains a novel epidermal growth factor-like domain that makes it unique among all pyridoxal-5'-phosphate-dependent enzymes.
- Department of Structural Biology and Crystallography, Institute of Molecular Biotechnology, Beutenbergstrasse 11, D-07745 Jena, Germany.
Organizational Affiliation: