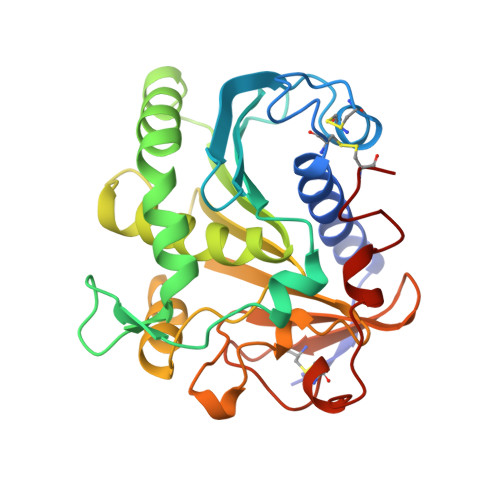
The crystal structure of lipase II from Rhizopus niveus at 2.2 A resolution.
Kohno, M., Funatsu, J., Mikami, B., Kugimiya, W., Matsuo, T., Morita, Y.(1996) J Biochem 120: 505-510
- PubMed: 8902613 Search on PubMed
- DOI: https://doi.org/10.1093/oxfordjournals.jbchem.a021442
- Primary Citation Related Structures:
1LGY - PubMed Abstract:
The crystal and molecular structure of Lipase II from Rhizopus niveus was analyzed using X-ray single crystal diffraction data at a resolution of 2.2 A. The structure was refined to an R-factor of 0.19 for all available data. This lipase was purified and crystallized as Lipase I, which contains two polypeptide chains combined through non-covalent interaction. However, during crystal growth, Lipase I was converted to Lipase II, which consists of a single polypeptide chain of 269 amino acid residues, by limited proteolysis. The structure of Lipase II shows a typical alpha/beta hydrolase fold containing the so-called nucleophilic elbow. The catalytic center of this enzyme is analogous to those of other neutral lipases and serine proteases. This catalytic center is sheltered by an alpha-helix lid, which appears in neutral lipases, opening the active site at the oil-water interface.
- Central Research Institute Tsukuba R&D Center, Fuji Oil Co., Ltd., Ibaraki. kouno@fujioil.co.jp
Organizational Affiliation: