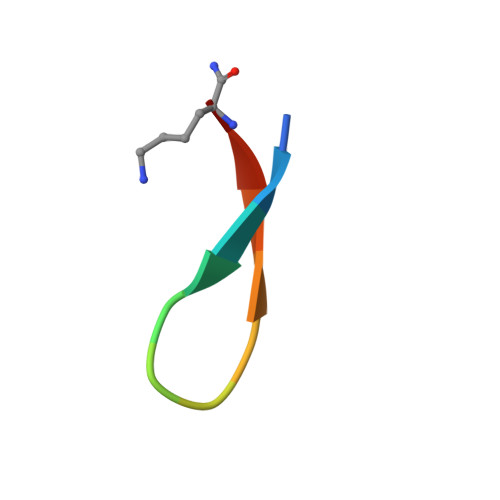
Tryptophan zippers: stable, monomeric beta -hairpins.
Cochran, A.G., Skelton, N.J., Starovasnik, M.A.(2001) Proc Natl Acad Sci U S A 98: 5578-5583
- PubMed: 11331745 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.091100898
- Primary Citation Related Structures:
1LE0, 1LE1, 1LE3 - PubMed Abstract:
A structural motif, the tryptophan zipper (trpzip), greatly stabilizes the beta-hairpin conformation in short peptides. Peptides (12 or 16 aa in length) with four different turn sequences are monomeric and fold cooperatively in water, as has been observed previously for some hairpin peptides. However, the folding free energies of the trpzips exceed substantially those of all previously reported beta-hairpins and even those of some larger designed proteins. NMR structures of three of the trpzip peptides reveal exceptionally well-defined beta-hairpin conformations stabilized by cross-strand pairs of indole rings. The trpzips are the smallest peptides to adopt an unique tertiary fold without requiring metal binding, unusual amino acids, or disulfide crosslinks.
- Department of Protein Engineering, Genentech, Inc., 1 DNA Way, South San Francisco, CA 94080, USA. andrea@gene.com
Organizational Affiliation: