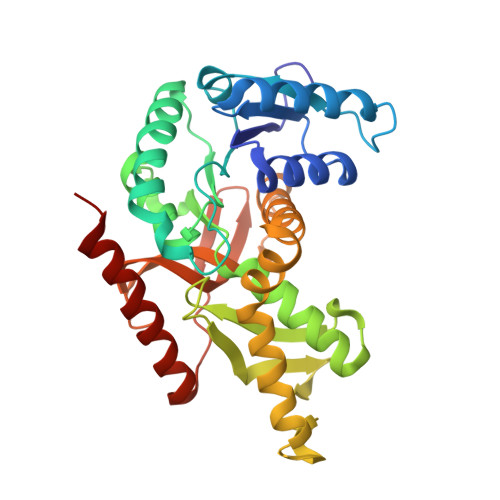
Structure of a ternary complex of an allosteric lactate dehydrogenase from Bacillus stearothermophilus at 2.5 A resolution.
Wigley, D.B., Gamblin, S.J., Turkenburg, J.P., Dodson, E.J., Piontek, K., Muirhead, H., Holbrook, J.J.(1992) J Mol Biology 223: 317-335
- PubMed: 1731077 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(92)90733-z
- Primary Citation Related Structures:
1LDN - PubMed Abstract:
We report the refined structure of a ternary complex of an allosterically activated lactate dehydrogenase, including the important active site loop. Eightfold non-crystallographic symmetry averaging was utilized to improve the density maps. Interactions between the protein and bound coenzyme and oxamate are described in relation to other studies using site-specific mutagenesis. Fructose 1,6-bisphosphate (FruP2) is bound to the enzyme across one of the 2-fold axes of the tetramer, with the two phosphate moieties interacting with two anion binding sites, one on each of two subunits, across this interface. However, because FruP2 binds at this special site, yet does not possess an internal 2-fold symmetry axis, the ligand is statistically disordered and binds to each site in two different orientations. Binding of FruP2 to the tetramer is signalled to the active site principally through two interactions with His188 and Arg173. His188 is connected to His195 (which binds the carbonyl group of the substrate) and Arg173 is connected to Arg171 (the residue that binds the carboxylate group of the substrate).
- Department of Biochemistry, University of Leicester, U.K.
Organizational Affiliation: