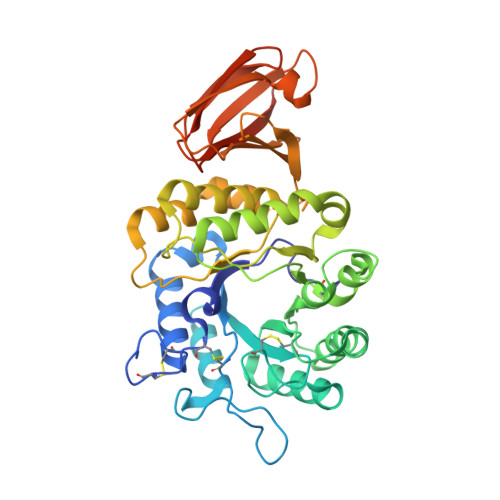
The 1.9 A structure of alpha-N-acetylgalactosaminidase: molecular basis of glycosidase deficiency diseases
Garman, S.C., Hannick, L., Zhu, A., Garboczi, D.N.(2002) Structure 10: 425-434
- PubMed: 12005440 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(02)00726-8
- Primary Citation Related Structures:
1KTB, 1KTC - PubMed Abstract:
In the lysosome, glycosidases degrade glycolipids, glycoproteins, and oligosaccharides. Mutations in glycosidases cause disorders characterized by the deposition of undegraded carbohydrates. Schindler and Fabry diseases are caused by the incomplete degradation of carbohydrates with terminal alpha-N-acetylgalactosamine and alpha-galactose, respectively. Here we present the X-ray structure of alpha-N-acetylgalactosaminidase (alpha-NAGAL), the glycosidase that removes alpha-N-acetylgalactosamine, and the structure with bound ligand. The active site residues of alpha-NAGAL are conserved in the closely related enzyme a-galactosidase A (alpha-GAL). The structure demonstrates the catalytic mechanisms of both enzymes and reveals the structural basis of mutations causing Schindler and Fabry diseases. As alpha-NAGAL and alpha-GAL produce type O "universal donor" blood from type A and type B blood, the alpha-NAGAL structure will aid in the engineering of improved enzymes for blood conversion.
- Structural Biology Section, Laboratory of Immunogenetics, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Rockville, Maryland 20852, USA. garman@alpha.niaid.nih.gov
Organizational Affiliation: