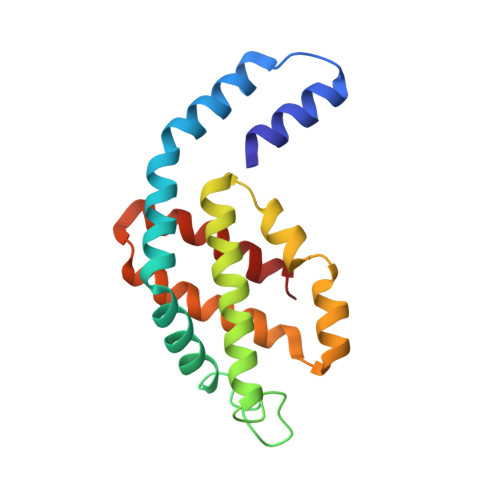
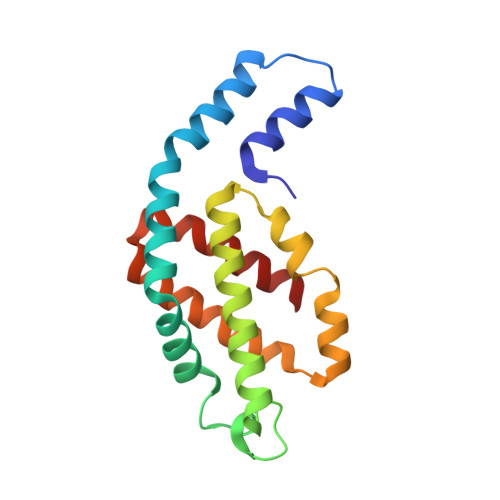
Crystal structure of Allophycocyanin from red algae Porphyra yezoensis at 2.2 A resolution
Liu, J.Y., Jiang, T., Zhang, J.P., Liang, D.C.(1999) J Biological Chem 274: 16945-16952
- PubMed: 10358042 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.274.24.16945
- Primary Citation Related Structures:
1KN1 - PubMed Abstract:
The crystal structure of allophycocyanin from red algae Porphyra yezoensis (APC-PY) at 2.2-A resolution has been determined by the molecular replacement method. The crystal belongs to space group R32 with cell parameters a = b = 105.3 A, c = 189.4 A, alpha = beta = 90 degrees, gamma = 120 degrees. After several cycles of refinement using program X-PLOR and model building based on the electron density map, the crystallographic R-factor converged to 19.3% (R-free factor is 26.9%) in the range of 10.0 to 2.2 A. The r.m.s. deviations of bond length and angles are 0.015 A and 2.9 degrees, respectively. In the crystal, two APC-PY trimers associate face to face into a hexamer. The assembly of two trimers within the hexamer is similar to that of C-phycocyanin (C-PC) and R-phycoerythrin (R-PE) hexamers, but the assembly tightness of the two trimers to the hexamer is not so high as that in C-PC and R-PE hexamers. The chromophore-protein interactions and possible pathway of energy transfer were discussed. Phycocyanobilin 1alpha84 of APC-PY forms 5 hydrogen bonds with 3 residues in subunit 2beta of another monomer. In R-PE and C-PC, chromophore 1alpha84 only forms 1 hydrogen bond with 2beta77 residue in subunit 2beta. This result may support and explain great spectrum difference exists between APC trimer and monomer.
- National Laboratory of Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, Chaoyang District, Beijing 100101, China.
Organizational Affiliation: