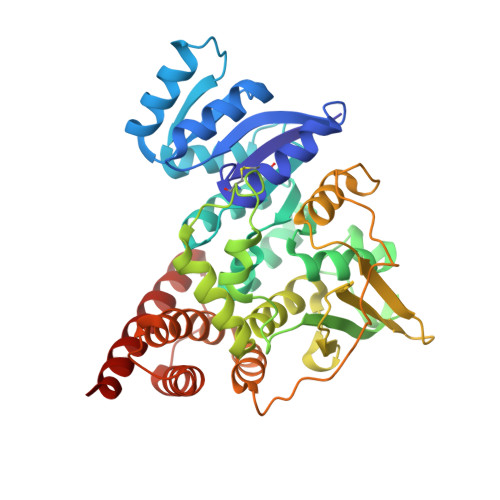
Crystal structure of 1-deoxy-D-xylulose-5-phosphate reductoisomerase, a crucial enzyme in the non-mevalonate pathway of isoprenoid biosynthesis.
Reuter, K., Sanderbrand, S., Jomaa, H., Wiesner, J., Steinbrecher, I., Beck, E., Hintz, M., Klebe, G., Stubbs, M.T.(2002) J Biological Chem 277: 5378-5384
- PubMed: 11741911 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M109500200
- Primary Citation Related Structures:
1K5H - PubMed Abstract:
We have solved the 2.5-A crystal structure of 1-deoxy-D-xylulose-5-phosphate reductoisomerase, an enzyme involved in the mevalonate-independent 2-C-methyl-D-erythritol-4-phosphate pathway of isoprenoid biosynthesis. The structure reveals that the enzyme is present as a homodimer. Each monomer displays a V-like shape and is composed of an amino-terminal dinucleotide binding domain, a connective domain, and a carboxyl-terminal four-helix bundle domain. The connective domain is responsible for dimerization and harbors most of the active site. The strictly conserved acidic residues Asp(150), Glu(152), Glu(231), and Glu(234) are clustered at the putative active site and are probably involved in the binding of divalent cations mandatory for enzyme activity. The connective and four-helix bundle domains show significant mobility upon superposition of the dinucleotide binding domains of the three conformational states present in the asymmetric unit of the crystal. A still more pronounced flexibility is observed for a loop spanning residues 186 to 216, which adopts two completely different conformations within the three protein conformers. A possible involvement of this loop in an induced fit during substrate binding is discussed.
- Institut für Pharmazeutische Chemie, Philipps-Universität, Marbacher Weg 6, D-35032 Marburg, Germany. reuterk@mailer.uni-marburg.de
Organizational Affiliation: