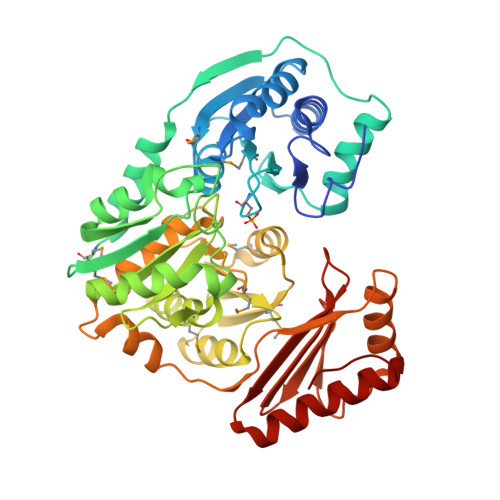
Crystal structure of PMM/PGM: an enzyme in the biosynthetic pathway of P. aeruginosa virulence factors.
Regni, C., Tipton, P.A., Beamer, L.J.(2002) Structure 10: 269-279
- PubMed: 11839312 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(02)00705-0
- Primary Citation Related Structures:
1K2Y, 1K35 - PubMed Abstract:
The enzyme phosphomannomutase/phosphoglucomutase (PMM/PGM) from P. aeruginosa is required for the biosynthesis of two bacterial exopolysaccharides: alginate and lipopolysaccharide (LPS). Both of these molecules play a role in the virulence of P. aeruginosa, an important human pathogen known for its ability to develop antibiotic resistance and cause chronic lung infections in cystic fibrosis patients. The crystal structure of PMM/PGM shows that the enzyme has four domains, three of which have a similar three-dimensional fold. Residues from all four domains of the protein contribute to the formation of a large active site cleft in the center of the molecule. Detailed information on the active site of PMM/PGM lays the foundation for structure-based inhibitor design. Inhibitors of sufficient potency and specificity should impair the biosynthesis of alginate and LPS, and may facilitate clearance of the bacteria by the host immune system and increase the efficacy of conventional antibiotic treatment against chronic P. aeruginosa infections.
- Department of Biochemistry, 117 Schweitzer Hall, University of Missouri-Columbia, Columbia, MO 65211, USA.
Organizational Affiliation: