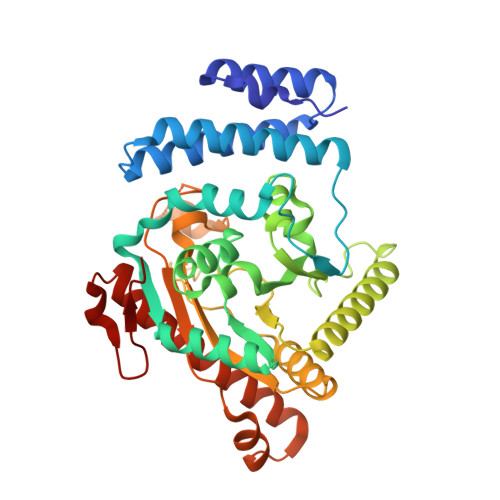
Analysis of the structure, substrate specificity, and mechanism of squash glycerol-3-phosphate (1)-acyltransferase.
Turnbull, A.P., Rafferty, J.B., Sedelnikova, S.E., Slabas, A.R., Schierer, T.P., Kroon, J.T., Simon, J.W., Fawcett, T., Nishida, I., Murata, N., Rice, D.W.(2001) Structure 9: 347-353
- PubMed: 11377195 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(01)00595-0
- Primary Citation Related Structures:
1K30 - PubMed Abstract:
Glycerol-3-phosphate (1)-acyltransferase(G3PAT) catalyzes the incorporation of an acyl group from either acyl-acyl carrier proteins (acylACPs) or acyl-CoAs into the sn-1 position of glycerol 3-phosphate to yield 1-acylglycerol-3-phosphate. G3PATs can either be selective, preferentially using the unsaturated fatty acid, oleate (C18:1), as the acyl donor, or nonselective, using either oleate or the saturated fatty acid, palmitate (C16:0), at comparable rates. The differential substrate specificity for saturated versus unsaturated fatty acids seen within this enzyme family has been implicated in the sensitivity of plants to chilling temperatures. The three-dimensional structure of recombinant G3PAT from squash chloroplast has been determined to 1.9 A resolution by X-ray crystallography using the technique of multiple isomorphous replacement and provides the first representative structure of an enzyme of this class. The tertiary structure of G3PAT comprises two domains, the larger of which, domain II, features an extensive cleft lined by hydrophobic residues and contains at one end a cluster of positively charged residues flanked by a H(X)(4)D motif, which is conserved amongst many glycerolipid acyltransferases. We predict that these hydrophobic and positively charged residues represent the binding sites for the fatty acyl substrate and the phosphate moiety of the glycerol 3-phosphate, respectively, and that the H(X)(4)D motif is a critical component of the enzyme's catalytic machinery.
- The Krebs Institute for Biomolecular Research, Department of Molecular Biology and Biotechnology, The University of Sheffield, S10 2TN, Sheffield, United Kingdom.
Organizational Affiliation: