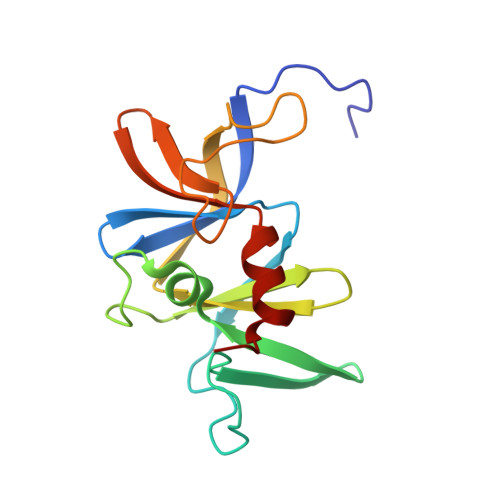
The solution structure and interactions of CheW from Thermotoga maritima.
Griswold, I.J., Zhou, H., Matison, M., Swanson, R.V., McIntosh, L.P., Simon, M.I., Dahlquist, F.W.(2002) Nat Struct Biol 9: 121-125
- PubMed: 11799399 Search on PubMed
- DOI: https://doi.org/10.1038/nsb753
- Primary Citation Related Structures:
1K0S - PubMed Abstract:
Using protein from the hyperthermophile Thermotoga maritima, we have determined the solution structure of CheW, an essential component in the formation of the bacterial chemotaxis signaling complex. The overall fold is similar to the regulatory domain of the chemotaxis kinase CheA. In addition, interactions of CheW with CheA were monitored by nuclear magnetic resonance (NMR) techniques. The chemical shift perturbation data show the probable contacts that CheW makes with CheA. In combination with previous genetic data, the structure also suggests a possible binding site for the chemotaxis receptor. These results provide a structural basis for a model in which CheW acts as a molecular bridge between CheA and the cytoplasmic tails of the receptor.
- Institute of Molecular Biology, Department of Chemistry, University of Oregon Eugene, Oregon 97403, USA.
Organizational Affiliation: