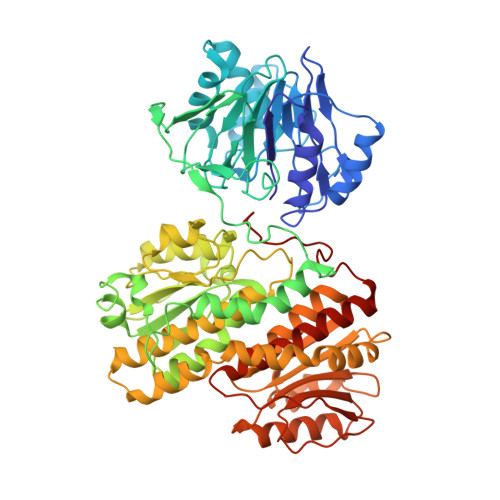
Channeling of ammonia in glucosamine-6-phosphate synthase.
Teplyakov, A., Obmolova, G., Badet, B., Badet-Denisot, M.A.(2001) J Mol Biology 313: 1093-1102
- PubMed: 11700065 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2001.5094
- Primary Citation Related Structures:
1JXA - PubMed Abstract:
Glucosamine-6-phosphate synthase catalyses the first and rate-limiting step in hexosamine metabolism, converting fructose 6-phosphate into glucosamine 6-phosphate in the presence of glutamine. The crystal structure of the Escherichia coli enzyme reveals the domain organisation of the homodimeric molecule. The 18 A hydrophobic channel sequestered from the solvent connects the glutaminase and isomerase active sites, and provides a means of ammonia transfer from glutamine to sugar phosphate. The C-terminal decapeptide sandwiched between the two domains plays a central role in the transfer. Based on the structure, a mechanism of enzyme action and self-regulation is proposed. It involves large domain movements triggered by substrate binding that lead to the formation of the channel.
- European Molecular Biology Laboratory, Notkestr. 85, D-22603 Hamburg, Germany. alexey@carb.inst.gov
Organizational Affiliation: