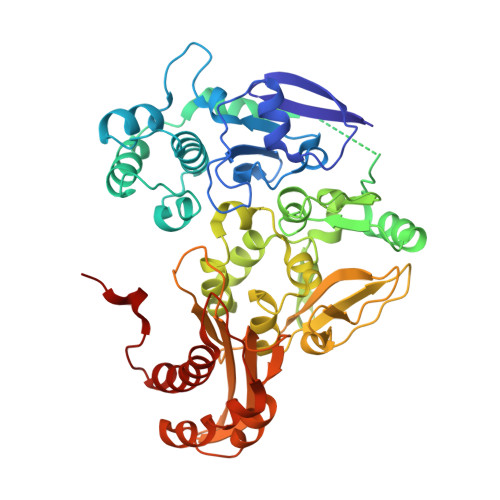
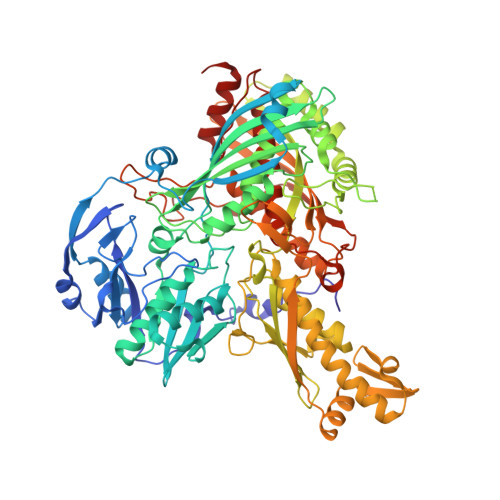
Crystal structures of the active and alloxanthine-inhibited forms of xanthine dehydrogenase from Rhodobacter capsulatus
Truglio, J.J., Theis, K., Leimkuhler, S., Rappa, R., Rajagopalan, K.V., Kisker, C.(2002) Structure 10: 115-125
- PubMed: 11796116 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(01)00697-9
- Primary Citation Related Structures:
1JRO, 1JRP - PubMed Abstract:
Xanthine dehydrogenase (XDH), a complex molybdo/iron-sulfur/flavoprotein, catalyzes the oxidation of hypoxanthine to xanthine followed by oxidation of xanthine to uric acid with concomitant reduction of NAD+. The 2.7 A resolution structure of Rhodobacter capsulatus XDH reveals that the bacterial and bovine XDH have highly similar folds despite differences in subunit composition. The NAD+ binding pocket of the bacterial XDH resembles that of the dehydrogenase form of the bovine enzyme rather than that of the oxidase form, which reduces O(2) instead of NAD+. The drug allopurinol is used to treat XDH-catalyzed uric acid build-up occurring in gout or during cancer chemotherapy. As a hypoxanthine analog, it is oxidized to alloxanthine, which cannot be further oxidized but acts as a tight binding inhibitor of XDH. The 3.0 A resolution structure of the XDH-alloxanthine complex shows direct coordination of alloxanthine to the molybdenum via a nitrogen atom. These results provide a starting point for the rational design of new XDH inhibitors.
- Department of Pharmacological Sciences, Center for Structural Biology, State University of New York at Stony Brook, Stony Brook, NY 11794, USA.
Organizational Affiliation: