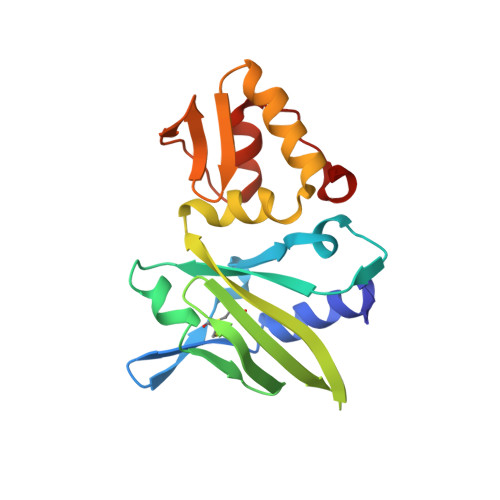
Solution structure of a viral DNA polymerase X and evidence for a mutagenic function.
Showalter, A.K., Byeon, I.J., Su, M.I., Tsai, M.D.(2001) Nat Struct Biol 8: 942-946
- PubMed: 11685239 Search on PubMed
- DOI: https://doi.org/10.1038/nsb1101-942
- Primary Citation Related Structures:
1JQR - PubMed Abstract:
The African swine fever virus DNA polymerase X (ASFV Pol X or Pol X), the smallest known nucleotide polymerase, has recently been reported to be an extremely low fidelity polymerase that may be involved in strategic mutagenesis of the viral genome. Here we report the solution structure of Pol X. The structure, unique within the realm of nucleotide polymerases, consists of only palm and fingers subdomains. Despite the absence of a thumb subdomain, which is important for DNA binding in other polymerases, we show that Pol X binds DNA with very high affinity. Further structural analyses suggest a novel mode of DNA binding that may contribute to low fidelity synthesis. We also demonstrate that the ASFV DNA ligase is a low fidelity ligase capable of sealing a nick that contains a G-G mismatch. This supports the hypothesis of a virus-encoded, mutagenic base excision repair pathway consisting of a tandem Pol X/ligase mutator.
- Department of Chemistry, The Ohio State University, Columbus Ohio 43210, USA.
Organizational Affiliation: