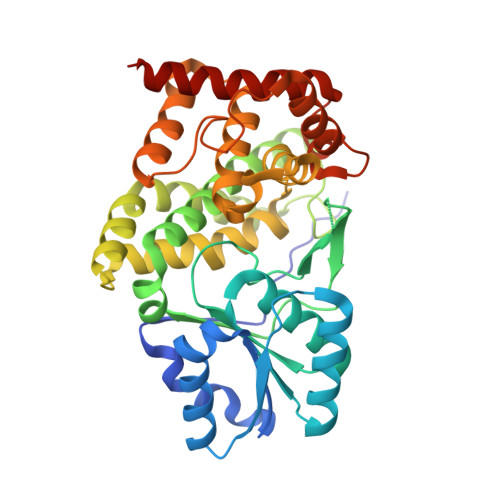
Glycerol dehydrogenase. structure, specificity, and mechanism of a family III polyol dehydrogenase.
Ruzheinikov, S.N., Burke, J., Sedelnikova, S., Baker, P.J., Taylor, R., Bullough, P.A., Muir, N.M., Gore, M.G., Rice, D.W.(2001) Structure 9: 789-802
- PubMed: 11566129 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(01)00645-1
- Primary Citation Related Structures:
1JPU, 1JQ5, 1JQA - PubMed Abstract:
Bacillus stearothermophilus glycerol dehydrogenase (GlyDH) (glycerol:NAD(+) 2-oxidoreductase, EC 1.1.1.6) catalyzes the oxidation of glycerol to dihydroxyacetone (1,3-dihydroxypropanone) with concomitant reduction of NAD(+) to NADH. Analysis of the sequence of this enzyme indicates that it is a member of the so-called iron-containing alcohol dehydrogenase family. Despite this sequence similarity, GlyDH shows a strict dependence on zinc for activity. On the basis of this, we propose to rename this group the family III metal-dependent polyol dehydrogenases. To date, no structural data have been reported for any enzyme in this group. The crystal structure of B. stearothermophilus glycerol dehydrogenase has been determined at 1.7 A resolution to provide structural insights into the mechanistic features of this family. The enzyme has 370 amino acid residues, has a molecular mass of 39.5 kDa, and is a homooctamer in solution. Analysis of the crystal structures of the free enzyme and of the binary complexes with NAD(+) and glycerol show that the active site of GlyDH lies in the cleft between the enzyme's two domains, with the catalytic zinc ion playing a role in stabilizing an alkoxide intermediate. In addition, the specificity of this enzyme for a range of diols can be understood, as both hydroxyls of the glycerol form ligands to the enzyme-bound Zn(2+) ion at the active site. The structure further reveals a previously unsuspected similarity to dehydroquinate synthase, an enzyme whose more complex chemistry shares a common chemical step with that catalyzed by glycerol dehydrogenase, providing a striking example of divergent evolution. Finally, the structure suggests that the NAD(+) binding domain of GlyDH may be related to that of the classical Rossmann fold by switching the sequence order of the two mononucleotide binding folds that make up this domain.
- Krebs Institute for Biomolecular Research, Department of Molecular Biology and Biotechnology, University of Sheffield, Firth Court, Western Bank, Sheffield S10 2TN, United Kingdom.
Organizational Affiliation: