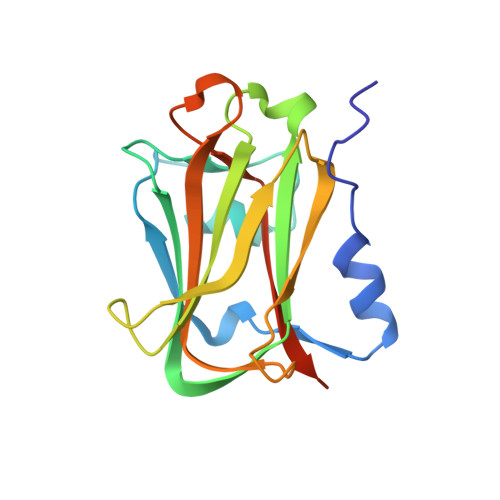
Crystal structure of the APC10/DOC1 subunit of the human anaphase-promoting complex
Wendt, K.S., Vodermaier, H.C., Jacob, U., Gieffers, C., Gmachl, M., Peters, J.M., Huber, R., Sondermann, P.(2001) Nat Struct Biol 8: 784-788
- PubMed: 11524682 Search on PubMed
- DOI: https://doi.org/10.1038/nsb0901-784
- Primary Citation Related Structures:
1JHJ - PubMed Abstract:
The anaphase-promoting complex (APC), or cyclosome, is a cell cycle-regulated ubiquitin ligase that controls progression through mitosis and the G1 phase of the cell cycle. The APC is composed of at least 11 subunits; no structure has been determined for any of these subunits. The subunit APC10/DOC1, a one-domain protein consisting of 185 amino acids, has a conserved core (residues 22-161) that is homologous to domains found in several other putative ubiquitin ligases and, therefore, may play a role in ubiquitination reactions. Here we report the crystal structure of human APC10 at 1.6 A resolution. The core of the protein is formed by a beta-sandwich that adopts a jellyroll fold. Unexpectedly, this structure is highly similar to ligand-binding domains of several bacterial and eukaryotic proteins, such as galactose oxidase and coagulation factor Va, raising the possibility that APC10 may function by binding a yet unidentified ligand. We further provide biochemical evidence that the C-terminus of APC10 binds to CDC27/APC3, an APC subunit that contains multiple tetratrico peptide repeats.
- Max-Planck-Institut für Biochemie, Abteilung Strukturforschung, Am Klopferspitz 18a, D-82152 Martinsried, Germany. wendt@biochem.mpg.de
Organizational Affiliation: