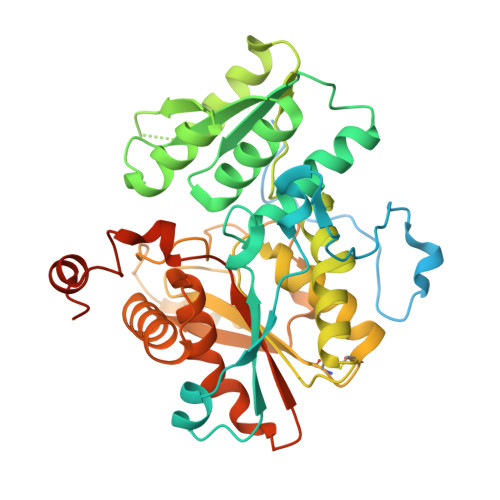
Structure of human cystathionine beta-synthase: a unique pyridoxal 5'-phosphate-dependent heme protein.
Meier, M., Janosik, M., Kery, V., Kraus, J.P., Burkhard, P.(2001) EMBO J 20: 3910-3916
- PubMed: 11483494 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/20.15.3910
- Primary Citation Related Structures:
1JBQ - PubMed Abstract:
Cystathionine beta-synthase (CBS) is a unique heme- containing enzyme that catalyzes a pyridoxal 5'-phosphate (PLP)-dependent condensation of serine and homocysteine to give cystathionine. Deficiency of CBS leads to homocystinuria, an inherited disease of sulfur metabolism characterized by increased levels of the toxic metabolite homocysteine. Here we present the X-ray crystal structure of a truncated form of the enzyme. CBS shares the same fold with O-acetylserine sulfhydrylase but it contains an additional N-terminal heme binding site. This heme binding motif together with a spatially adjacent oxidoreductase active site motif could explain the regulation of its enzyme activity by redox changes.
- M.E.Müller Institute for Structural Biology, Biozentrum, University of Basel, Klingelbergstrasse 70, CH-4056 Basel, Switzerland.
Organizational Affiliation: