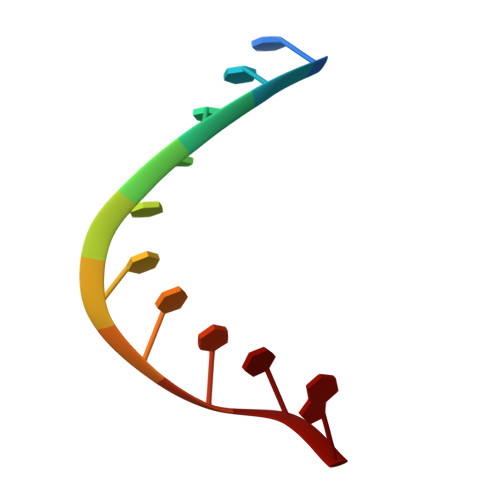
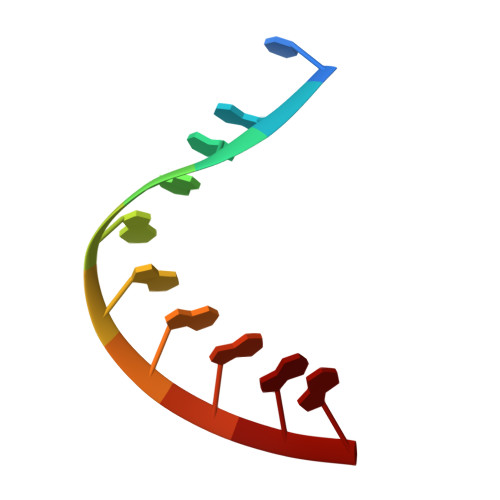
Crystal structure of an RNADNA hybrid reveals intermolecular intercalation: Dimer formation by base-pair swapping
Han, G.W., Kopka, M.L., Langs, D., Sawaya, M.R., Dickerson, R.E.(2003) Proc Natl Acad Sci U S A 100: 9214-9219
- PubMed: 12872000 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1533326100
- Primary Citation Related Structures:
1JB8 - PubMed Abstract:
An intermolecular intercalation of base pairs was found at the CA step in the I222 crystal structure of the RNA.DNA hybrid, r(CAAAGAAAAG).d(CTTTTCTTTG), which contains two-thirds of the polypurine tract sequence of HIV-1 with a substitution of cytosine for the initial adenine. This sequence crystallized in both P212121 and I222 space groups, with an rms difference of only 0.63 A between residues 3 to 18 of the two forms. P212121 and I222 helices are both A-like, but intercalation occurs only in the I222 crystal form. The present structure shows bases stacked in parallel rather than perpendicular as in intercalated DNA (I-DNA). The base intercalation is also different from zipper-like meshing of bases seen in the center of the crystal structure of d(GCGAAAGCT), which does not have Watson-Crick base pairing. The base-step intercalation seen here is reminiscent of domain swapping in proteins; therefore, we call this phenomenon "base-pair swapping." It involves a highly mobile CA step and seems to be sequence-specific and electrostatically stable without disrupting Watson-Crick interactions. It also exhibits a large rise concurrent with unwinding of the helix (low twist). We present a base-pair swapping dimer in nucleic acids.
- Molecular Biology Institute, University of California, Los Angeles, CA 90095-1570, USA.
Organizational Affiliation: