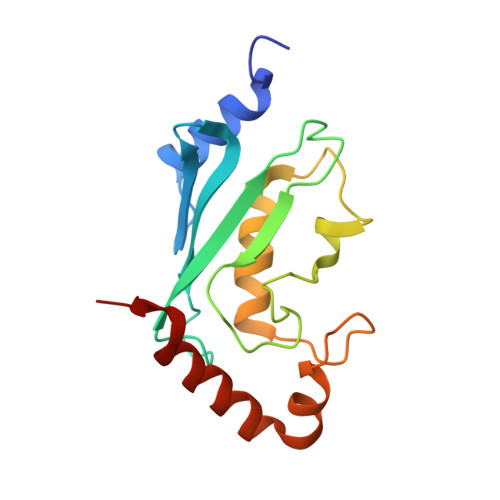
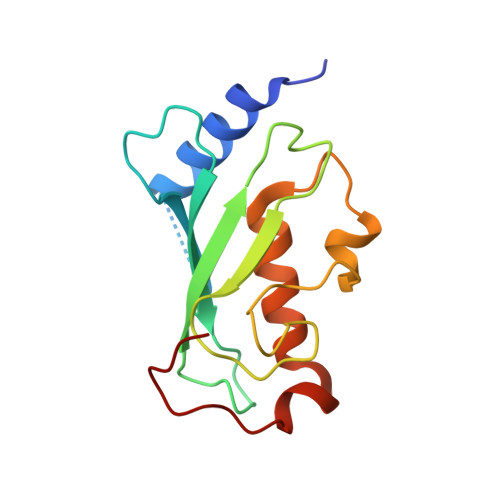
Molecular insights into polyubiquitin chain assembly: crystal structure of the Mms2/Ubc13 heterodimer.
VanDemark, A.P., Hofmann, R.M., Tsui, C., Pickart, C.M., Wolberger, C.(2001) Cell 105: 711-720
- PubMed: 11440714 Search on PubMed
- DOI: https://doi.org/10.1016/s0092-8674(01)00387-7
- Primary Citation Related Structures:
1JAT, 1JBB - PubMed Abstract:
While the signaling properties of ubiquitin depend on the topology of polyubiquitin chains, little is known concerning the molecular basis of specificity in chain assembly and recognition. UEV/Ubc complexes have been implicated in the assembly of Lys63-linked polyubiquitin chains that act as a novel signal in postreplicative DNA repair and I kappa B alpha kinase activation. The crystal structure of the Mms2/Ubc13 heterodimer shows the active site of Ubc13 at the intersection of two channels that are potential binding sites for the two substrate ubiquitins. Mutations that destabilize the heterodimer interface confer a marked UV sensitivity, providing direct evidence that the intact heterodimer is necessary for DNA repair. Selective mutations in the channels suggest a molecular model for specificity in the assembly of Lys63-linked polyubiquitin signals.
- Department of Biophysics and Biophysical Chemistry, School of Medicine, Johns Hopkins University, Baltimore, MD, USA.
Organizational Affiliation: