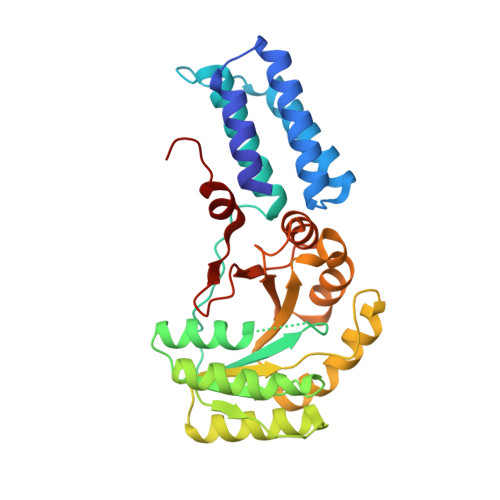
The crystal structure of the conserved GTPase of SRP54 from the archaeon Acidianus ambivalens and its comparison with related structures suggests a model for the SRP-SRP receptor complex.
Montoya, G., Kaat, K., Moll, R., Schafer, G., Sinning, I.(2000) Structure 8: 515-525
- PubMed: 10801496 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(00)00131-3
- Primary Citation Related Structures:
1J8M, 1J8Y - PubMed Abstract:
Protein targeting to the endoplasmic reticulum in eukaryotes and to the cell membrane in prokaryotes is mediated by the signal recognition particle (SRP) and its receptor (SR). Both contain conserved GTPase domains in the signal-peptide-binding proteins (SRP54 and Ffh) and the SR proteins (SRalpha and FtsY). These GTPases are involved in the regulation of protein targeting. Most studies so far have focussed on the SRP machinery of mammals and bacteria, leaving the SRP system of archaea less well understood. We report the crystal structure of the conserved GTPase (NG-Ffh) from the thermophilic archaeon Acidianus ambivalens at 2.0 A resolution and of the Thr112-->Ala mutant, which is inactive in GTP hydrolysis. This is the first structure of an SRP component from an archaeon and allows for a detailed comparison with related structures from Escherichia coli and thermophilic bacteria. In particular, differences in the conserved consensus regions for nucleotide binding and the subdomain interfaces are observed, which provide information about the regulation of the GTPase. These interactions allow us to propose a common signalling mechanism for the SRP-SR system. The overall structure of SRP-GTPases is well conserved between bacteria and archaea, which indicates strong similarities in the regulation of the SRP-targeting pathway. Surprisingly, structure comparisons identified a homodimeric ATP-binding protein as the closest relative. A heterodimer model for the SRP-SR interaction is presented.
- European Molecular Biology Laboratory, Structural Biology Programme, Heidelberg, D-69017, Germany.
Organizational Affiliation: