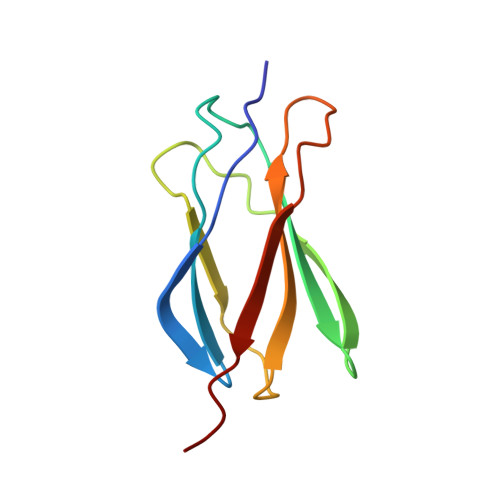
NMR structure of human fibronectin EDA.
Niimi, T., Osawa, M., Yamaji, N., Yasunaga, K., Sakashita, H., Mase, T., Tanaka, A., Fujita, S.(2001) J Biomol NMR 21: 281-284
- PubMed: 11775745 Search on PubMed
- DOI: https://doi.org/10.1023/a:1012947209393
- Primary Citation Related Structures:
1J8K