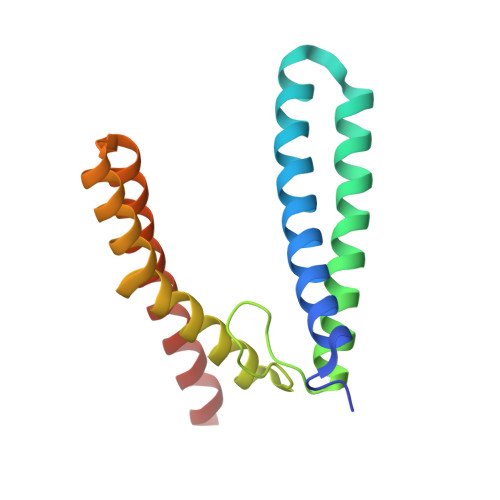
The Crystal Structure of Sulerythrin, A Rubrerythrin-like Protein from A Strictly Aerobic Archaeon, Sulfolobus tokodaii strain 7, shows unexpected domain swapping
Fushinobu, S., Shoun, H., Wakagi, T.(2003) Biochemistry 42: 11707-11715
- PubMed: 14529281 Search on PubMed
- DOI: https://doi.org/10.1021/bi034220b
- Primary Citation Related Structures:
1J30 - PubMed Abstract:
Sulerythrin is the first rubrerythrin-like protein to be isolated from an aerobic organism, Sulfolobus tokodaii strain 7, and it lacks a C-terminal rubredoxin-like FeS(4) domain. The protein purified from Sulfolobus cells was crystallized, and the crystal structure was determined at 1.7 A resolution. The dimer of sulerythrin exhibited "domain-swapping" at the loop connecting alphaB and alphaC, hybrid four-helix bundles consisting of alphaA/B and alphaC/D being formed. The structure and atomic identity of the binuclear metal center were determined by means of anomalous scattering analysis. The site contained 1.0 mol of hexacoordinate Fe, 0.80-0.87 mol of tetracoordinate Zn, and 0.73-0.88 mol of putative O(2) per monomer. The metal ions were found at exchanged positions compared to those in the Fe/Zn-containing rubrerythrin from Desulfovibrio vulgaris. The results demonstrate that the binuclear metal center of rubrerythrin-like proteins is plastic in its ability to bind metal ions.
- Department of Biotechnology, The University of Tokyo, 1-1-1 Yayoi, Bunkyo-ku, Tokyo 113-8657, Japan. asfushi@mail.ecc.u-tokyo.ac.jp
Organizational Affiliation: