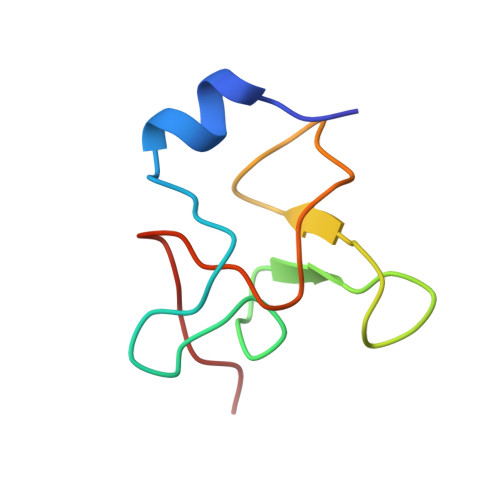
Three-dimensional structure of the high-potential iron-sulfur protein isolated from the purple phototrophic bacterium Rhodocyclus tenuis determined and refined at 1.5 A resolution.
Rayment, I., Wesenberg, G., Meyer, T.E., Cusanovich, M.A., Holden, H.M.(1992) J Mol Biology 228: 672-686
- PubMed: 1453470 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(92)90849-f
- Primary Citation Related Structures:
1ISU - PubMed Abstract:
The molecular structure of the high-potential iron-sulfur protein (HiPIP) isolated from the phototrophic bacterium, Rhodocyclus tenuis, has been solved and refined to a nominal resolution of 1.5 A with a crystallographic R-factor of 17.3% for all measured X-ray data from 30 A to 1.5 A. It is the smallest of the HiPIP structures studied thus far with 62 amino acid residues. Crystals used in the investigation belonged to the space group P2(1) with unit cell dimensions of a = 36.7 A, b = 52.6 A, c = 27.6 A and beta = 90.8 degrees and contained two molecules per asymmetric unit. The structure was solved by a combination of multiple isomorphous replacement with two heavy-atom derivatives, anomalous scattering from the iron-sulfur cluster, symmetry averaging and solvent flattening. The folding motif for this HiPIP is characterized by one small alpha-helix, six Type I turns, an approximate Type II turn and one Type I' turn. As in other HiPIPs, the iron-sulfur cluster is co-ordinated by four cysteinyl ligands and exhibits a cubane-like motif. These cysteinyl ligands are all located in Type I turns. The hydrogen bonding around the metal cluster in the R. tenuis protein is similar to the patterns observed in the Chromatium vinosum and Ectothiorhodospira halophila HiPIPs. Several of the amino acid residues invariant in the previously determined C. vinosum and E. halophila structures are not retained in the R. tenuis molecule. There are 13 solvent molecules structurally conserved between the two R. tenuis HiPIP molecules in the asymmetric unit, some of which are important for stabilizing surface loops. Interestingly, while it is assumed that this HiPIP functions as a monomer in solution, the two molecules in the asymmetric unit pack as a dimer and are related to each other by an approximate twofold rotation axis.
- Institute for Enzyme Research, Graduate School, College of Agricultural and Life Sciences, University of Wisconsin, Madison 53705.
Organizational Affiliation: