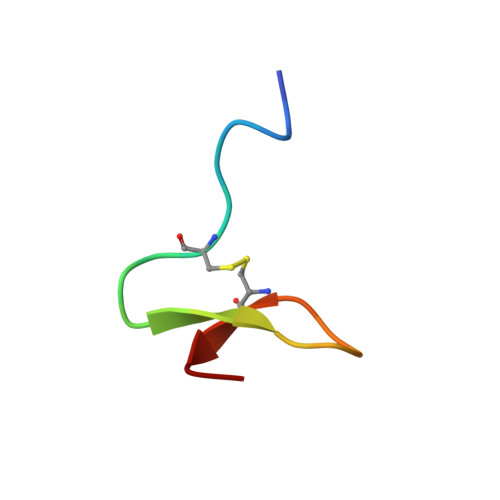
Solution structure of paralytic peptide of silkworm, Bombyx mori
Miura, K., Kamimura, M., Aizawa, T., Kiuchi, M., Hayakawa, Y., Mizuguchi, M., Kawano, K.(2002) Peptides 23: 2111-2116
- PubMed: 12535689 Search on PubMed
- DOI: https://doi.org/10.1016/s0196-9781(02)00254-1
- Primary Citation Related Structures:
1IRR - PubMed Abstract:
Paralytic peptide of Bombyx mori (BmPP) is one of the multifunctional ENF-peptides; the name of "ENF" is the consensus N-terminal amino acid sequence of the family peptides. We revealed that BmPP significantly possesses growth-blocking activity and plasmatocyte-spreading activity and that its activity profiles are different from those of another ENF-family peptide, namely, the growth-blocking peptide of Pseudaletia separata (PsGBP). We also determined the NMR structures of BmPP and PsGBP under the same conditions, which revealed the structural differences of the first and second beta-turn regions between the two peptides. On the basis of our results, it can be considered that the tertiary structural difference in these peptides may cause their different profiles of growth-blocking activity.
- Bio-oriented Technology Research Advancement Institution, 1-40-2 Nisshin, Saitama, Saitama 331-8537, Japan.
Organizational Affiliation: